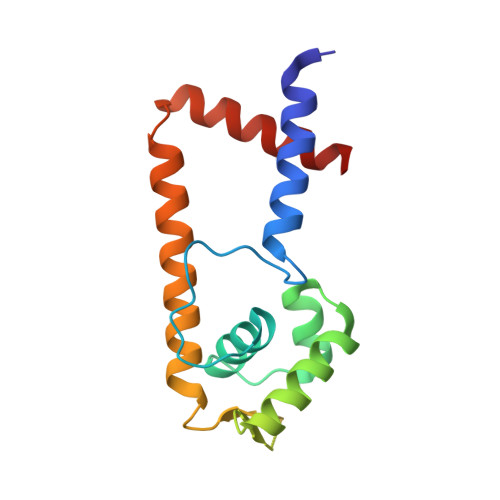
Crystal Structure of the Zinc-Dependent MarR Family Transcriptional Regulator AdcR in the Zn(II)-Bound State.
Guerra, A.J., Dann, C.E., Giedroc, D.P.(2011) J Am Chem Soc 133: 19614-19617
- PubMed: 22085181
- DOI: https://doi.org/10.1021/ja2080532
- Primary Citation Related Structures:
3TGN - PubMed Abstract:
Streptococcus pneumoniae adhesin competence regulator (AdcR), the first metal-dependent member of the multiple antibiotic resistance regulator (MarR) family of proteins, represses the transcription of a high-affinity zinc-specific uptake transporter, a group of surface antigen zinc-binding pneumococcal histidine triad proteins (PhtA, PhtB, PhtD, and PhtE), and an AdcA homologue (AdcAII). The 2.0 Å resolution structure of Zn(II)-bound AdcR reveals a highly helical two-fold-symmetric dimer with two distinct metal-binding sites per protomer. Zn(II) is tetrahedrally coordinated by E24, H42, H108, and H112 in what defines the primary sensing site in AdcR. Site 2 is a tetracoordinate site whose function is currently unknown. NMR methyl group perturbation experiments reveal that Zn(II) drives a global change in the structure of apo-AdcR that stabilizes a conformation that is compatible with DNA binding. This co-repression mechanism is unprecedented in MarR transcriptional regulators.
- Department of Chemistry, Indiana University, Bloomington, Indiana 47405, USA.
Organizational Affiliation: