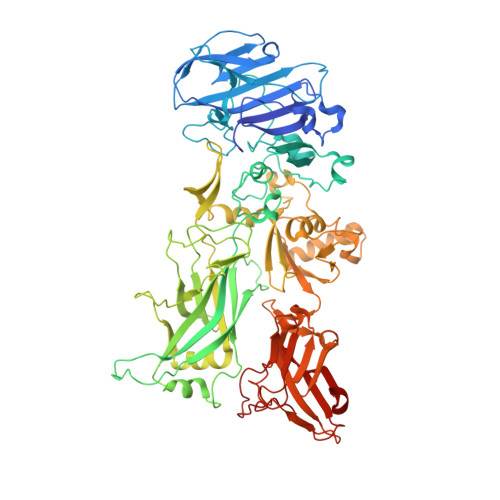
Domain flexibility modulates the heterogeneous assembly mechanism of anthrax toxin protective antigen.
Feld, G.K., Kintzer, A.F., Tang, I.I., Thoren, K.L., Krantz, B.A.(2012) J Mol Biology 415: 159-174
- PubMed: 22063095 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.10.035
- Primary Citation Related Structures:
3TEW, 3TEX, 3TEY, 3TEZ - PubMed Abstract:
The three protein components of anthrax toxin are nontoxic individually, but they form active holotoxin complexes upon assembly. The role of the protective antigen (PA) component of the toxin is to deliver two other enzyme components, lethal factor and edema factor, across the plasma membrane and into the cytoplasm of target cells. PA is produced as a proprotein, which must be proteolytically activated; generally, cell surface activation is mediated by a furin family protease. Activated PA can then assemble into one of two noninterconverting oligomers, a homoheptamer and a homooctamer, which have unique properties. Herein we describe molecular determinants that influence the stoichiometry of PA in toxin complexes. By tethering PA domain 4 (D4) to domain 2 with two different-length cross-links, we can control the relative proportions of PA heptamers and octamers. The longer cross-link favors octamer formation, whereas the shorter one favors formation of the heptamer. X-ray crystal structures of PA (up to 1.45 Å resolution), including these cross-linked PA constructs, reveal that a hinge-like movement of D4 correlates with the relative preference for each oligomeric architecture. Furthermore, we report the conformation of the flexible loop containing the furin cleavage site and show that, for efficient processing, the furin site cannot be moved ~5 or 6 residues within the loop. We propose that there are different orientations of D4 relative to the main body of PA that favor the formation of either the heptamer or the octamer.
- Department of Chemistry, University of California, Berkeley, CA 94720, USA.
Organizational Affiliation: