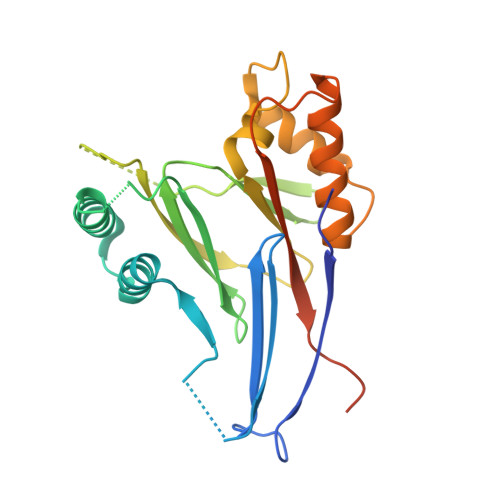
Structure of the phosphatase domain of the cell fate determinant SpoIIE from Bacillus subtilis.
Levdikov, V.M., Blagova, E.V., Rawlings, A.E., Jameson, K., Tunaley, J., Hart, D.J., Barak, I., Wilkinson, A.J.(2012) J Mol Biology 415: 343-358
- PubMed: 22115775 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.11.017
- Primary Citation Related Structures:
3T91, 3T9Q - PubMed Abstract:
Sporulation in Bacillus subtilis begins with an asymmetric cell division producing two genetically identical cells with different fates. SpoIIE is a membrane protein that localizes to the polar cell division sites where it causes FtsZ to relocate from mid-cell to form polar Z-rings. Following polar septation, SpoIIE establishes compartment-specific gene expression in the smaller forespore cell by dephosphorylating the anti-sigma factor antagonist SpoIIAA, leading to the release of the RNA polymerase sigma factor σ(F) from an inhibitory complex with the anti-sigma factor SpoIIAB. SpoIIE therefore couples morphological development to differential gene expression. Here, we determined the crystal structure of the phosphatase domain of SpoIIE to 2.6 Å spacing, revealing a domain-swapped dimer. SEC-MALLS (size-exclusion chromatography with multi-angle laser light scattering) analysis however suggested a monomer as the principal form in solution. A model for the monomer was derived from the domain-swapped dimer in which 2 five-stranded β-sheets are packed against one another and flanked by α-helices in an αββα arrangement reminiscent of other PP2C-type phosphatases. A flap region that controls access of substrates to the active site in other PP2C phosphatases is diminished in SpoIIE, and this observation correlates with the presence of a single manganese ion in the active site of SpoIIE in contrast to the two or three metal ions present in other PP2C enzymes. Mapping of a catalogue of mutational data onto the structure shows a clustering of sites whose point mutation interferes with the proper coupling of asymmetric septum formation to sigma factor activation and identifies a surface involved in intramolecular signaling.
- Structural Biology Laboratory, Department of Chemistry, University of York, York YO10 5YW, UK.
Organizational Affiliation: