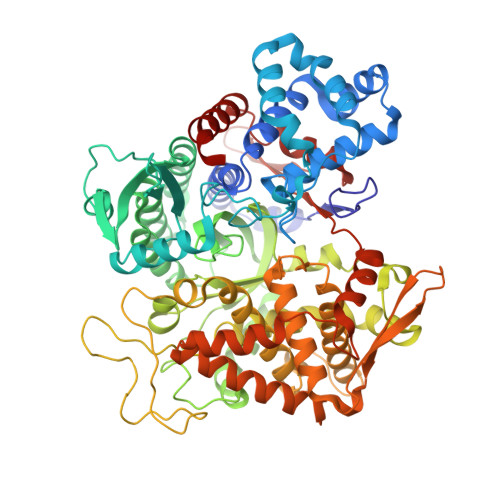
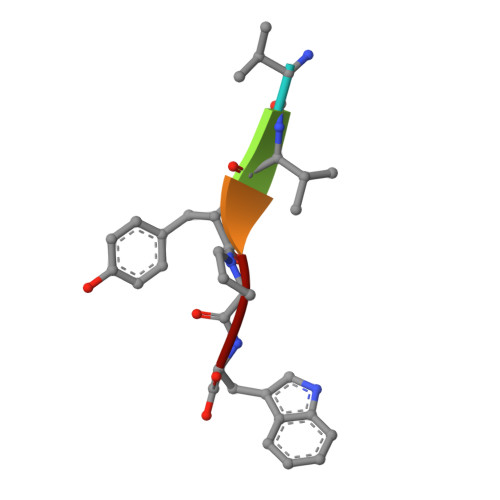
Entropy-driven binding of opioid peptides induces a large domain motion in human dipeptidyl peptidase III
Bezerra, G.A., Dobrovetsky, E., Viertlmayr, R., Dong, A., Binter, A., Abramic, M., Macheroux, P., Dhe-Paganon, S., Gruber, K.(2012) Proc Natl Acad Sci U S A 109: 6525-6530
- PubMed: 22493238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1118005109
- Primary Citation Related Structures:
3FVY, 3T6B, 3T6J - PubMed Abstract:
Opioid peptides are involved in various essential physiological processes, most notably nociception. Dipeptidyl peptidase III (DPP III) is one of the most important enkephalin-degrading enzymes associated with the mammalian pain modulatory system. Here we describe the X-ray structures of human DPP III and its complex with the opioid peptide tynorphin, which rationalize the enzyme's substrate specificity and reveal an exceptionally large domain motion upon ligand binding. Microcalorimetric analyses point at an entropy-dominated process, with the release of water molecules from the binding cleft ("entropy reservoir") as the major thermodynamic driving force. Our results provide the basis for the design of specific inhibitors that enable the elucidation of the exact role of DPP III and the exploration of its potential as a target of pain intervention strategies.
- Institute of Molecular Biosciences, University of Graz, A-8010 Graz, Austria.
Organizational Affiliation: