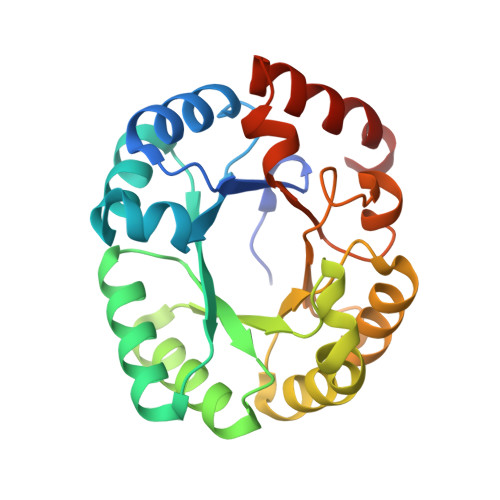
Crystal structure of the mutant S127A of orotidine 5'-monophosphate decarboxylase from Methanobacterium thermoautotrophicum complexed with the inhibitor 6azaUMP
Fedorov, A.A., Fedorov, E.V., Desai, B., Iiams, V., Gerlt, J.A., Almo, S.C.To be published.