Design of HIV-1 Protease Inhibitors with C3-Substituted Hexahydrocyclopentafuranyl Urethanes as P2-Ligands: Synthesis, Biological Evaluation, and Protein-Ligand X-ray Crystal Structure.
Ghosh, A.K., Chapsal, B.D., Parham, G.L., Steffey, M., Agniswamy, J., Wang, Y.F., Amano, M., Weber, I.T., Mitsuya, H.(2011) J Med Chem 54: 5890-5901
- PubMed: 21800876 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm200649p
- Primary Citation Related Structures:
3ST5 - PubMed Abstract:
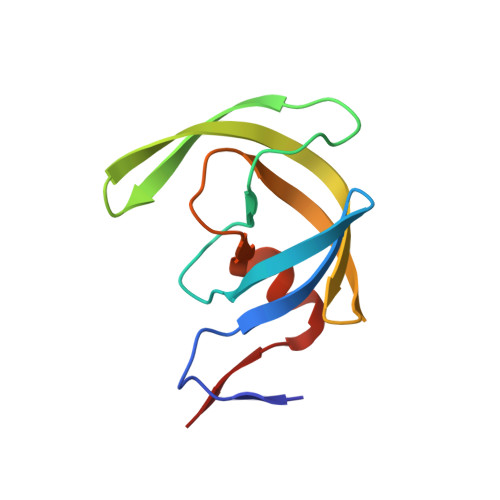
We report the design, synthesis, biological evaluation, and the X-ray crystal structure of a novel inhibitor bound to the HIV-1 protease. Various C3-functionalized cyclopentanyltetrahydrofurans (Cp-THF) were designed to interact with the flap Gly48 carbonyl or amide NH in the S2-subsite of the HIV-1 protease. We investigated the potential of those functionalized ligands in combination with hydroxyethylsulfonamide isosteres. Inhibitor 26 containing a 3-(R)-hydroxyl group on the Cp-THF core displayed the most potent enzyme inhibitory and antiviral activity. Our studies revealed a preference for the 3-(R)-configuration over the corresponding 3-(S)-derivative. Inhibitor 26 exhibited potent activity against a panel of multidrug-resistant HIV-1 variants. A high resolution X-ray structure of 26-bound HIV-1 protease revealed important molecular insight into the ligand-binding site interactions.
- Department of Chemistry, Purdue University, West Lafayette, Indiana 47907, USA. akghosh@purdue.edu
Organizational Affiliation: