High-Throughput Genetic Identification of Functionally Important Regions of the Yeast DEAD-Box Protein Mss116p.
Mohr, G., Del Campo, M., Turner, K.G., Gilman, B., Wolf, R.Z., Lambowitz, A.M.(2011) J Mol Biology 413: 952-972
- PubMed: 21945532
- DOI: https://doi.org/10.1016/j.jmb.2011.09.015
- Primary Citation Related Structures:
3SQW, 3SQX - PubMed Abstract:
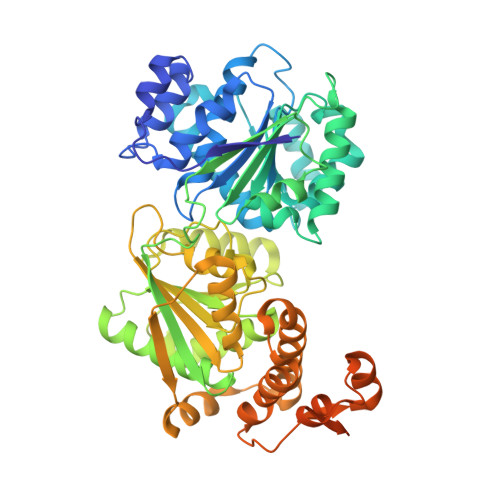
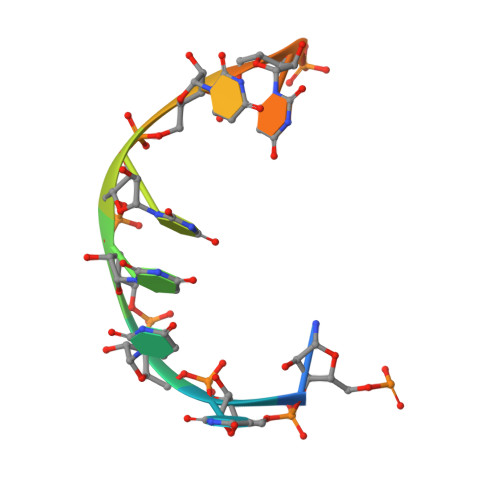
The Saccharomyces cerevisiae DEAD-box protein Mss116p is a general RNA chaperone that functions in splicing mitochondrial group I and group II introns. Recent X-ray crystal structures of Mss116p in complex with ATP analogs and single-stranded RNA show that the helicase core induces a bend in the bound RNA, as in other DEAD-box proteins, while a C-terminal extension (CTE) induces a second bend, resulting in RNA crimping. Here, we illuminate these structures by using high-throughput genetic selections, unigenic evolution, and analyses of in vivo splicing activity to comprehensively identify functionally important regions and permissible amino acid substitutions throughout Mss116p. The functionally important regions include those containing conserved sequence motifs involved in ATP and RNA binding or interdomain interactions, as well as previously unidentified regions, including surface loops that may function in protein-protein interactions. The genetic selections recapitulate major features of the conserved helicase motifs seen in other DEAD-box proteins but also show surprising variations, including multiple novel variants of motif III (SAT). Patterns of amino acid substitutions indicate that the RNA bend induced by the helicase core depends on ionic and hydrogen-bonding interactions with the bound RNA; identify a subset of critically interacting residues; and indicate that the bend induced by the CTE results primarily from a steric block. Finally, we identified two conserved regions-one the previously noted post II region in the helicase core and the other in the CTE-that may help displace or sequester the opposite RNA strand during RNA unwinding.
- Institute for Cellular and Molecular Biology, Department of Chemistry and Biochemistry, and Section of Molecular Genetics and Microbiology, School of Biological Sciences, University of Texas at Austin, Austin, TX 78712, USA.
Organizational Affiliation: