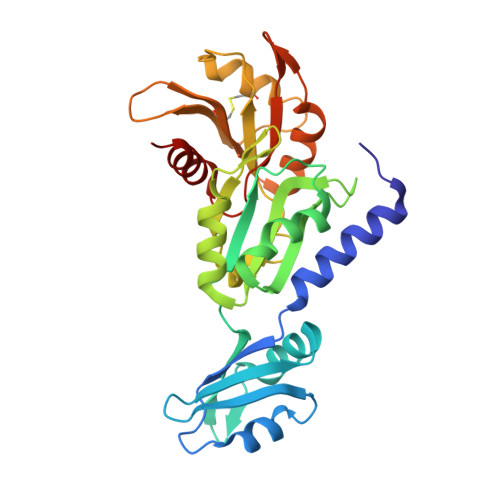
Apo raver1 structure reveals distinct RRM domain orientations.
Rangarajan, E.S., Lee, J.H., Izard, T.(2011) Protein Sci 20: 1464-1470
- PubMed: 21633983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.664
- Primary Citation Related Structures:
3SMZ - PubMed Abstract:
Raver1 is a multifunctional protein that modulates both alternative splicing and focal adhesion assembly by binding to the nucleoplasmic splicing repressor polypyrimidine tract protein (PTB) or to the cytoskeletal proteins vinculin and α-actinin. The amino-terminal region of raver1 has three RNA recognition motif (RRM1, RRM2, and RRM3) domains, and RRM1 interacts with the vinculin tail (Vt) domain and vinculin mRNA. We previously determined the crystal structure of the raver1 RRM1-3 domains in complex with Vt at 2.75 Å resolution. Here, we report crystal structure of the unbound raver1 RRM1-3 domains at 2 Å resolution. The apo structure reveals that a bound sulfate ion disrupts an electrostatic interaction between the RRM1 and RRM2 domains, triggering a large relative domain movement of over 30°. Superposition with other RNA-bound RRM structures places the sulfate ion near the superposed RNA phosphate group suggesting that this is the raver1 RNA binding site. While several single and some tandem RRM domain structures have been described, to the best of our knowledge, this is the second report of a three-tandem RRM domain structure.
- Cell Adhesion Laboratory, Department of Cancer Biology, The Scripps Research Institute, Jupiter, Florida 33458, USA.
Organizational Affiliation: