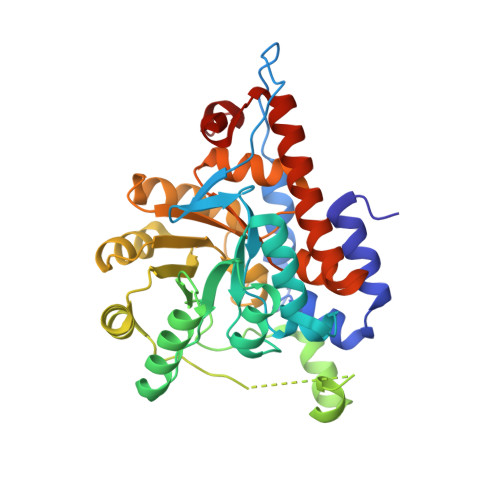
High resolution crystal structure of rat long chain hydroxy acid oxidase in complex with the inhibitor 4-carboxy-5-[(4-chlorophenyl)sulfanyl]-1, 2, 3-thiadiazole. Implications for inhibitor specificity and drug design.
Chen, Z.W., Vignaud, C., Jaafar, A., Levy, B., Gueritte, F., Guenard, D., Lederer, F., Mathews, F.S.(2012) Biochimie 94: 1172-1179
- PubMed: 22342614 Search on PubMed
- DOI: https://doi.org/10.1016/j.biochi.2012.02.003
- Primary Citation Related Structures:
3SGZ - PubMed Abstract:
Long chain hydroxy acid oxidase (LCHAO) is responsible for the formation of methylguanidine, a toxic compound with elevated serum levels in patients with chronic renal failure. Its isozyme glycolate oxidase (GOX), has a role in the formation of oxalate, which can lead to pathological deposits of calcium oxalate, in particular in the disease primary hyperoxaluria. Inhibitors of these two enzymes may have therapeutic value. These enzymes are the only human members of the family of FMN-dependent l-2-hydroxy acid-oxidizing enzymes, with yeast flavocytochrome b(2) (Fcb2) among its well studied members. We screened a chemical library for inhibitors, using in parallel rat LCHAO, human GOX and the Fcb2 flavodehydrogenase domain (FDH). Among the hits was an inhibitor, CCPST, with an IC(50) in the micromolar range for all three enzymes. We report here the crystal structure of a complex between this compound and LCHAO at 1.3 Å resolution. In comparison with a lower resolution structure of this enzyme, binding of the inhibitor induces a conformational change in part of the TIM barrel loop 4, as well as protonation of the active site histidine. The CCPST interactions are compared with those it forms with human GOX and those formed by two other inhibitors with human GOX and spinach GOX. These compounds differ from CCPST in having the sulfur replaced with a nitrogen in the five-membered ring as well as different hydrophobic substituents. The possible reason for the ∼100-fold difference in affinity between these two series of inhibitors is discussed. The present results indicate that specificity is an issue in the quest for therapeutic inhibitors of either LCHAO or GOX, but they may give leads for this quest.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, 660 South Euclid Avenue, St Louis, MO 63110, USA.
Organizational Affiliation: