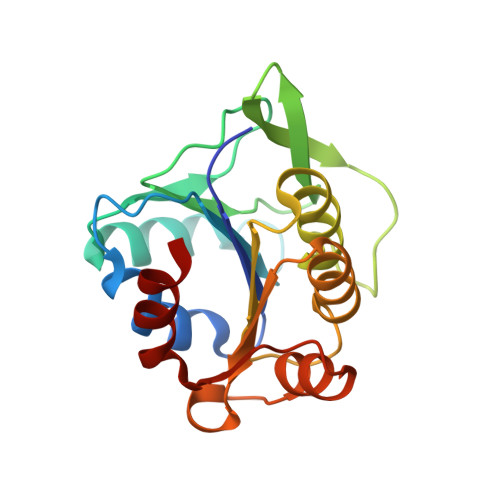
An insight into the interaction mode between CheB and chemoreceptor from two crystal structures of CheB methylesterase catalytic domain
Cho, K.H., Crane, B.R., Park, S.Y.(2011) Biochem Biophys Res Commun 411: 69-75
- PubMed: 21722627 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2011.06.090
- Primary Citation Related Structures:
3SFT - PubMed Abstract:
We have determined 2.2 Å resolution crystal structure of Thermotoga maritima CheB methylesterase domain to provide insight into the interaction mode between CheB and chemoreceptors. T. maritima CheB methylesterase domain has identical topology of a modified doubly-wound α/β fold that was observed from the previously reported Salmonella typhimurium counterpart, but the analysis of the electrostatic potential surface near the catalytic triad indicated considerable charge distribution difference. As the CheB demethylation consensus sites of the chemoreceptors, the CheB substrate, are not uniquely conserved between T. maritima and S. typhimurium, such surfaces with differing electrostatic properties may reflect CheB regions that mediate protein-protein interaction. Via the computational docking of the two T. maritima and S. typhimurium CheB structures to the respective T. maritima and Escherichia coli chemoreceptors, we propose a CheB:chemoreceptor interaction mode.
- School of Systems Biomedical Science, Soongsil University, Seoul, Republic of Korea.
Organizational Affiliation: