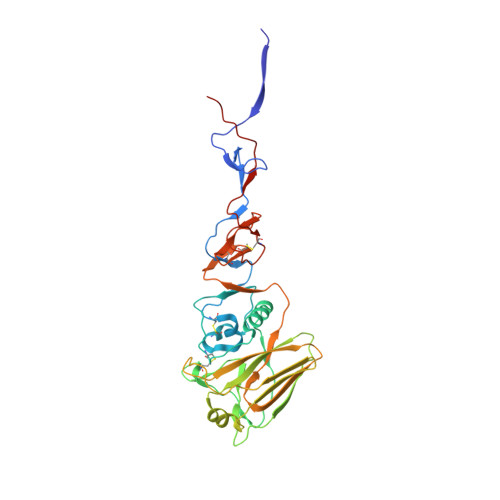
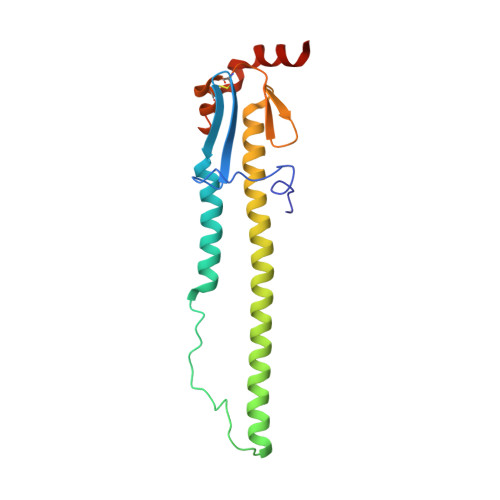
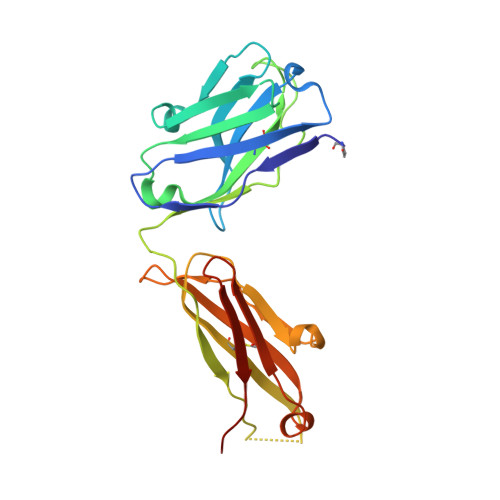
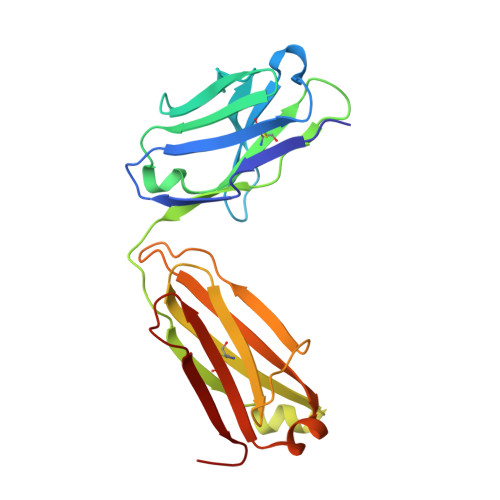
A highly conserved neutralizing epitope on group 2 influenza A viruses.
Ekiert, D.C., Friesen, R.H., Bhabha, G., Kwaks, T., Jongeneelen, M., Yu, W., Ophorst, C., Cox, F., Korse, H.J., Brandenburg, B., Vogels, R., Brakenhoff, J.P., Kompier, R., Koldijk, M.H., Cornelissen, L.A., Poon, L.L., Peiris, M., Koudstaal, W., Wilson, I.A., Goudsmit, J.(2011) Science 333: 843-850
- PubMed: 21737702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1204839
- Primary Citation Related Structures:
3SDY - PubMed Abstract:
Current flu vaccines provide only limited coverage against seasonal strains of influenza viruses. The identification of V(H)1-69 antibodies that broadly neutralize almost all influenza A group 1 viruses constituted a breakthrough in the influenza field. Here, we report the isolation and characterization of a human monoclonal antibody CR8020 with broad neutralizing activity against most group 2 viruses, including H3N2 and H7N7, which cause severe human infection. The crystal structure of Fab CR8020 with the 1968 pandemic H3 hemagglutinin (HA) reveals a highly conserved epitope in the HA stalk distinct from the epitope recognized by the V(H)1-69 group 1 antibodies. Thus, a cocktail of two antibodies may be sufficient to neutralize most influenza A subtypes and, hence, enable development of a universal flu vaccine and broad-spectrum antibody therapies.
- Department of Molecular Biology, Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: