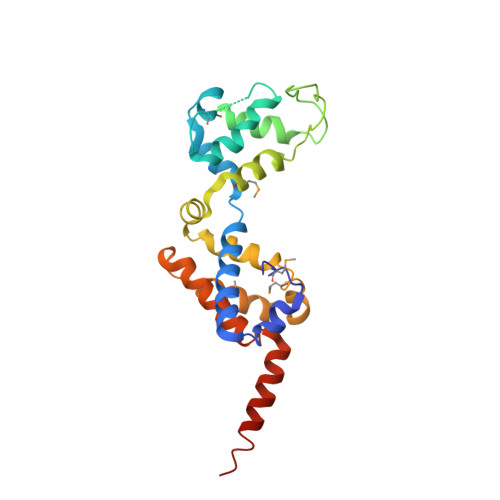
Mechanism of actin filament nucleation by Vibrio VopL and implications for tandem W domain nucleation.
Namgoong, S., Boczkowska, M., Glista, M.J., Winkelman, J.D., Rebowski, G., Kovar, D.R., Dominguez, R.(2011) Nat Struct Mol Biol 18: 1060-1067
- PubMed: 21873985 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2109
- Primary Citation Related Structures:
3RYL - PubMed Abstract:
Pathogen proteins targeting the actin cytoskeleton often serve as model systems to understand their more complex eukaryotic analogs. We show that the strong actin filament nucleation activity of Vibrio parahaemolyticus VopL depends on its three W domains and on its dimerization through a unique VopL C-terminal domain (VCD). The VCD shows a previously unknown all-helical fold and interacts with the pointed end of the actin nucleus, contributing to the nucleation activity directly and through duplication of the W domain repeat. VopL promotes rapid cycles of filament nucleation and detachment but generally has no effect on elongation. Profilin inhibits VopL-induced nucleation by competing for actin binding to the W domains. Combined, the results suggest that VopL stabilizes a hexameric double-stranded pointed end nucleus. Analysis of hybrid constructs of VopL and the eukaryotic nucleator Spire suggest that Spire may also function as a dimer in cells.
- Department of Physiology, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Organizational Affiliation: