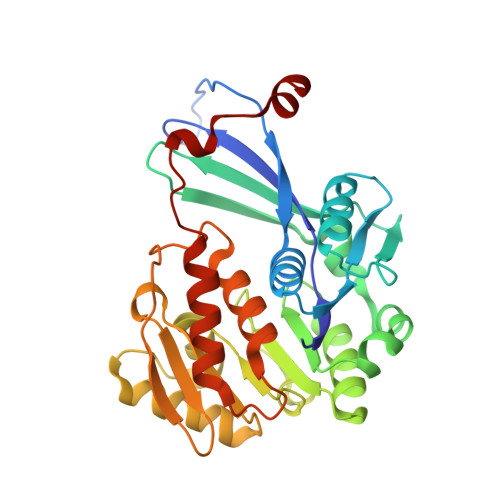
Crystal structure of Sa239 reveals the structural basis for the activation of ribokinase by monovalent cations.
Li, J., Wang, C., Wu, Y., Wu, M., Wang, L., Wang, Y., Zang, J.(2012) J Struct Biol 177: 578-582
- PubMed: 22198595
- DOI: https://doi.org/10.1016/j.jsb.2011.12.010
- Primary Citation of Related Structures:
3RY7 - PubMed Abstract:
Ribokinase is responsible for catalyzing the reaction of d-ribose and ATP to produce ribose-5-phosphate and ADP, which can be activated by monovalent cations such as potassium, cesium and ammonium. However, the exact activation mechanism of ribokinase remains elusive. Here we report the crystal structure of Sa239, a ribokinase from Staphylococcus aureus, in the absence of monovalent ions. In addition to the dimer form similar to that observed in Escherichia coli ribokinase structure, the structure of Sa239 demonstrates that the C-terminal tail protrudes from the remaining part and interacts with the neighboring molecule, resulting in an unexpected dimerization form. By comparing the structure of Sa239 to E. coli ribokinase, we propose that binding of the monovalent cation triggers the conformational change of the large ATP loop to organize the formation of nucleotide binding pocket, thus enabling ATP binding and enhancing catalytic activity. Our study uncovers the detailed structural basis for the activation mechanism of ribokinase by monovalent cations.
- Hefei National Laboratory for Physical Sciences at Microscale, School of Life Sciences, University of Science and Technology of China, 96 Jinzhai Road, Hefei, Anhui 230026, People's Republic of China.
Organizational Affiliation: