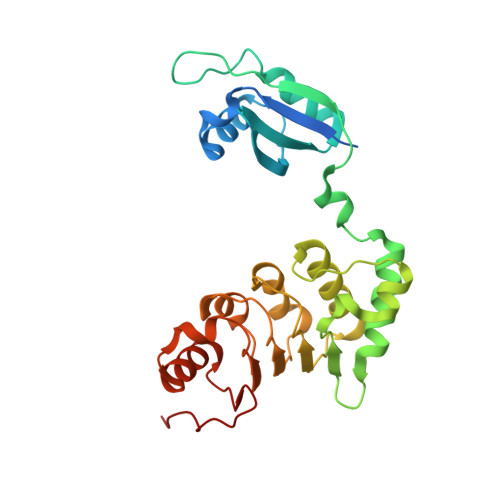
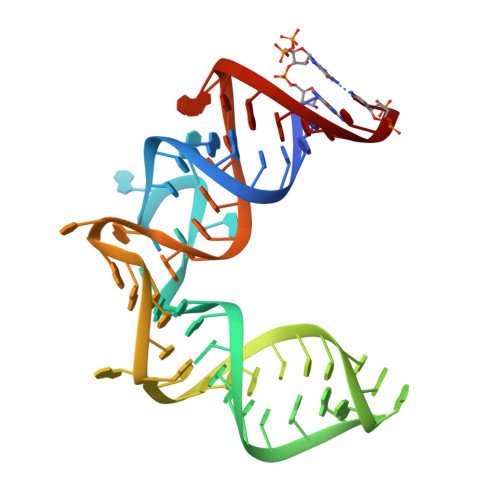
Structure-function studies of nucleocytoplasmic transport of retroviral genomic RNA by mRNA export factor TAP.
Teplova, M., Wohlbold, L., Khin, N.W., Izaurralde, E., Patel, D.J.(2011) Nat Struct Mol Biol 18: 990-998
- PubMed: 21822283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2094
- Primary Citation Related Structures:
3RW6, 3RW7 - PubMed Abstract:
mRNA export is mediated by the TAP-p15 heterodimer, which belongs to the family of NTF2-like export receptors. TAP-p15 heterodimers also bind to the constitutive transport element (CTE) present in simian type D retroviral RNAs, and they mediate the export of viral unspliced RNAs to the host cytoplasm. We have solved the crystal structure of the RNA recognition and leucine-rich repeat motifs of TAP bound to one symmetrical half of the CTE RNA. L-shaped conformations of protein and RNA are involved in a mutual molecular embrace on complex formation. We have monitored the impact of structure-guided mutations on binding affinities in vitro and transport assays in vivo. Our studies define the principles by which CTE RNA subverts the mRNA export receptor TAP, thereby facilitating the nuclear export of viral genomic RNAs, and, more generally, provide insights on cargo RNA recognition by mRNA export receptors.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York, USA.
Organizational Affiliation: