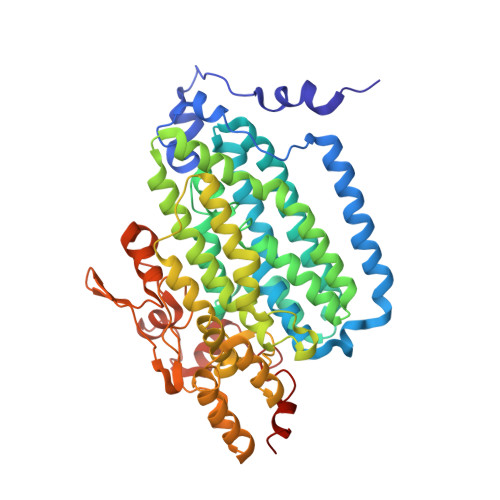
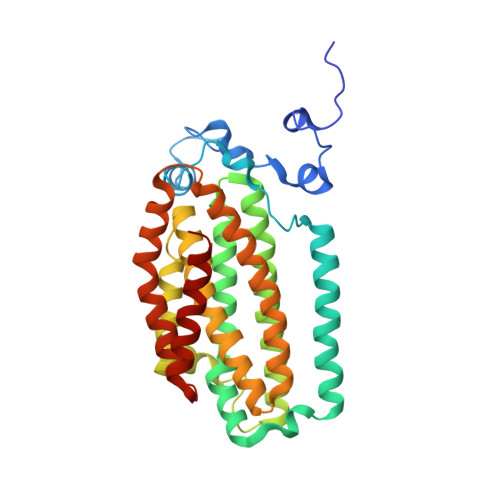
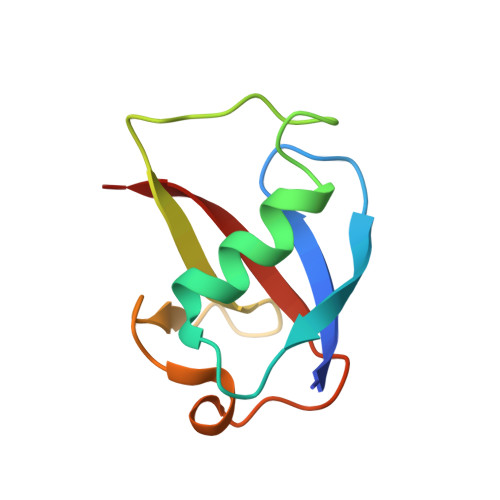
Tracking a defined route for O2 migration in a dioxygen-activating diiron enzyme.
Song, W.J., Gucinski, G., Sazinsky, M.H., Lippard, S.J.(2011) Proc Natl Acad Sci U S A 108: 14795-14800
- PubMed: 21859951 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1106514108
- Primary Citation Related Structures:
3RN9, 3RNA, 3RNB, 3RNC, 3RNE, 3RNF, 3RNG - PubMed Abstract:
For numerous enzymes reactive toward small gaseous compounds, growing evidence indicates that these substrates diffuse into active site pockets through defined pathways in the protein matrix. Toluene/o-xylene monooxygenase hydroxylase is a dioxygen-activating enzyme. Structural analysis suggests two possible pathways for dioxygen access through the α-subunit to the diiron center: a channel or a series of hydrophobic cavities. To distinguish which is utilized as the O(2) migration pathway, the dimensions of the cavities and the channel were independently varied by site-directed mutagenesis and confirmed by X-ray crystallography. The rate constants for dioxygen access to the diiron center were derived from the formation rates of a peroxodiiron(III) intermediate, generated upon treatment of the diiron(II) enzyme with O(2). This reaction depends on the concentration of dioxygen to the first order. Altering the dimensions of the cavities, but not the channel, changed the rate of dioxygen reactivity with the enzyme. These results strongly suggest that voids comprising the cavities in toluene/o-xylene monooxygenase hydroxylase are not artifacts of protein packing/folding, but rather programmed routes for dioxygen migration through the protein matrix. Because the cavities are not fully connected into the diiron active center in the enzyme resting state, conformational changes will be required to facilitate dioxygen access to the diiron center. We propose that such temporary opening and closing of the cavities may occur in all bacterial multicomponent monooxygenases to control O(2) consumption for efficient catalysis. Our findings suggest that other gas-utilizing enzymes may employ similar structural features to effect substrate passage through a protein matrix.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, MA 02139, USA.
Organizational Affiliation: