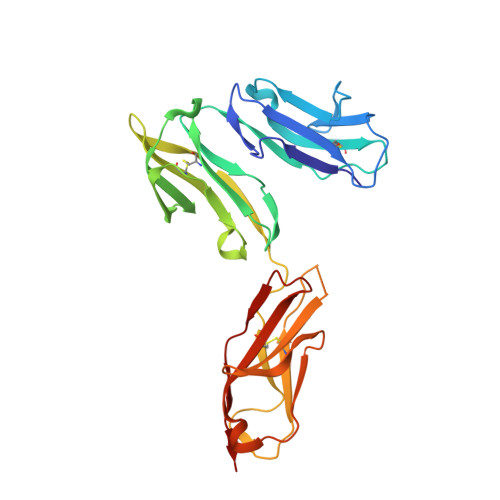Crystal structure of Fc gamma receptor I and its implication in high affinity gamma-immunoglobulin binding.
Lu, J., Ellsworth, J.L., Hamacher, N., Oak, S.W., Sun, P.D.(2011) J Biological Chem 286: 40608-40613
- PubMed: 21965667 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.257550
- Primary Citation Related Structures:
3RJD - PubMed Abstract:
Fcγ receptors (FcγRs) play critical roles in humoral and cellular immune responses through interactions with the Fc region of immunoglobulin G (IgG). Among them, FcγRI is the only high affinity receptor for IgG and thus is a potential target for immunotherapy. Here we report the first crystal structure of an FcγRI with all three extracellular Ig-like domains (designated as D1, D2, and D3). The structure shows that, first, FcγRI has an acute D1-D2 hinge angle similar to that of FcεRI but much smaller than those observed in the low affinity Fcγ receptors. Second, the D3 domain of FcγRI is positioned away from the putative IgG binding site on the receptor and is thus unlikely to make direct contacts with Fc. Third, the replacement of FcγRIII FG-loop ((171)LVGSKNV(177)) with that of FcγRI ((171)MGKHRY(176)) resulted in a 15-fold increase in IgG(1) binding affinity, whereas a valine insertion in the FcγRI FG-loop ((171)MVGKHRY(177)) abolished the affinity enhancement. Thus, the FcγRI FG-loop with its conserved one-residue deletion is critical to the high affinity IgG binding. The structural results support FcγRI binding to IgG in a similar mode as its low affinity counterparts. Taken together, our study suggests a molecular mechanism for the high affinity IgG recognition by FcγRI and provides a structural basis for understanding its physiological function and its therapeutic implication in treating autoimmune diseases.
- Structural Immunology Section, Laboratory of Immunogenetics, NIAID, National Institutes of Health, Rockville, Maryland, USA.
Organizational Affiliation: