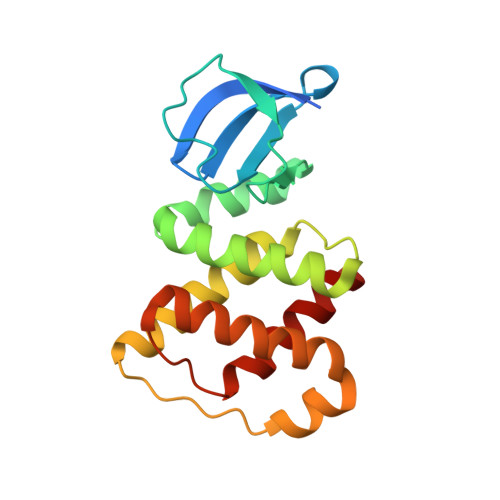
Structure of the RBD-PRDI fragment of the antiterminator protein GlcT.
Himmel, S., Grosse, C., Wolff, S., Schwiegk, C., Becker, S.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 751-756
- PubMed: 22750856 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112020635
- Primary Citation Related Structures:
3RIO - PubMed Abstract:
GlcT is a transcriptional antiterminator protein that is involved in regulation of glucose metabolism in Bacillus subtilis. Antiterminator proteins bind specific RNA sequences, thus preventing the formation of overlapping terminator stem-loops. The structure of a fragment (residues 3-170) comprising the RNA-binding domain (RBD) and the first regulatory domain (PRDI) of GlcT was solved at 2.0 Å resolution with one molecule in the asymmetric unit. The two domains are connected by a helical linker. Their interface is mostly constituted by hydrophobic interactions.
- Department of NMR-based Structural Biology, Max Planck Institute for Biophysical Chemistry, Göttingen, Germany.
Organizational Affiliation: