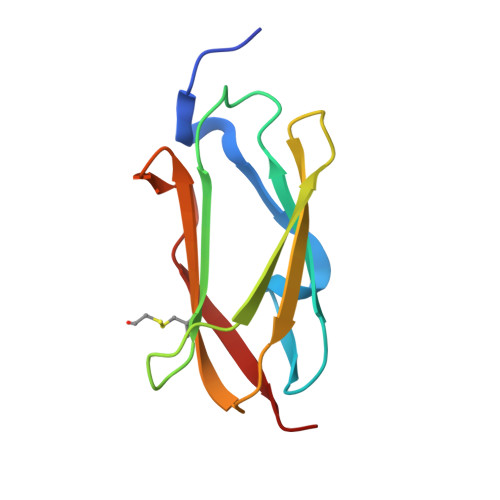
Structure of filamin A immunoglobulin-like repeat 10 from Homo sapiens.
Page, R.C., Clark, J.G., Misra, S.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 871-876
- PubMed: 21821884 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111024249
- Primary Citation Related Structures:
3RGH - PubMed Abstract:
Filamin A (FlnA) plays a critical role in cytoskeletal organization, cell motility and cellular signaling. FlnA utilizes different binding sites on a series of 24 immunoglobulin-like domains (Ig repeats) to interact with diverse cytosolic proteins and with cytoplasmic portions of membrane proteins. Mutations in a specific domain, Ig10 (FlnA-Ig10), are correlated with two severe forms of the otopalatodigital syndrome spectrum disorders Melnick-Needles syndrome and frontometaphyseal dysplasia. The crystal structure of FlnA-Ig10 determined at 2.44 Å resolution provides insight into the perturbations caused by these mutations.
- Department of Molecular Cardiology, Lerner Research Institute, Cleveland Clinic, Cleveland, OH 44195, USA. pager2@ccf.org
Organizational Affiliation: