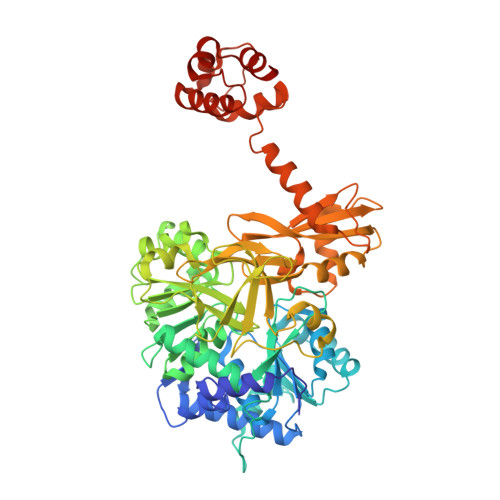
Structural and Functional Investigation of the Intermolecular Interaction between NRPS Adenylation and Carrier Protein Domains.
Sundlov, J.A., Shi, C., Wilson, D.J., Aldrich, C.C., Gulick, A.M.(2012) Chem Biol 19: 188-198
- PubMed: 22365602 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2011.11.013
- Primary Citation Related Structures:
3RG2 - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) are modular proteins that produce peptide antibiotics and siderophores. These enzymes act as catalytic assembly lines where substrates, covalently bound to integrated carrier domains, are delivered to adjacent catalytic domains. The carrier domains are initially loaded by adenylation domains, which use two distinct conformations to catalyze sequentially the adenylation of the substrate and the thioesterification of the pantetheine cofactor. We have used a mechanism-based inhibitor to determine the crystal structure of an engineered adenylation-carrier domain protein illustrating the intermolecular interaction between the adenylation and carrier domains. This structure enabled directed mutations to improve the interaction between nonnative partner proteins. Comparison with prior NRPS adenylation domain structures provides insights into the assembly line dynamics of these modular enzymes.
- Hauptman-Woodward Institute and Department of Structural Biology, University at Buffalo, Buffalo, NY 14203 USA.
Organizational Affiliation: