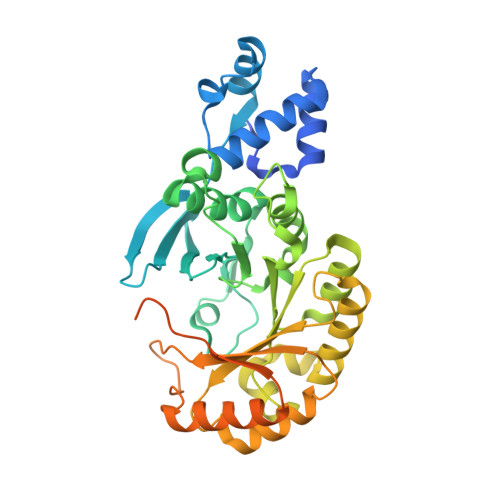
Structural basis for methyl transfer by a radical SAM enzyme.
Boal, A.K., Grove, T.L., McLaughlin, M.I., Yennawar, N.H., Booker, S.J., Rosenzweig, A.C.(2011) Science 332: 1089-1092
- PubMed: 21527678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1205358
- Primary Citation Related Structures:
3RF9, 3RFA - PubMed Abstract:
The radical S-adenosyl-L-methionine (SAM) enzymes RlmN and Cfr methylate 23S ribosomal RNA, modifying the C2 or C8 position of adenosine 2503. The methyl groups are installed by a two-step sequence involving initial methylation of a conserved Cys residue (RlmN Cys(355)) by SAM. Methyl transfer to the substrate requires reductive cleavage of a second equivalent of SAM. Crystal structures of RlmN and RlmN with SAM show that a single molecule of SAM coordinates the [4Fe-4S] cluster. Residue Cys(355) is S-methylated and located proximal to the SAM methyl group, suggesting the SAM that is involved in the initial methyl transfer binds at the same site. Thus, RlmN accomplishes its complex reaction with structural economy, harnessing the two most important reactivities of SAM within a single site.
- Department of Molecular Biosciences, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: