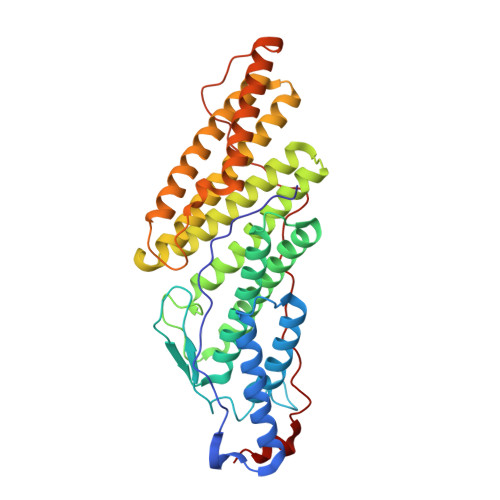
The Phe105 Loop of Alix Bro1 Domain Plays a Key Role in HIV-1 Release.
Sette, P., Mu, R., Dussupt, V., Jiang, J., Snyder, G., Smith, P., Xiao, T.S., Bouamr, F.(2011) Structure 19: 1485-1495
- PubMed: 21889351
- DOI: https://doi.org/10.1016/j.str.2011.07.016
- Primary Citation of Related Structures:
3R9M, 3RAU - PubMed Abstract:
Alix and cellular paralogs HD-PTP and Brox contain N-terminal Bro1 domains that bind ESCRT-III CHMP4. In contrast to HD-PTP and Brox, expression of the Bro1 domain of Alix alleviates HIV-1 release defects that result from interrupted access to ESCRT. In an attempt to elucidate this functional discrepancy, we solved the crystal structures of the Bro1 domains of HD-PTP and Brox. They revealed typical "boomerang" folds they share with the Bro1 Alix domain. However, they each contain unique structural features that may be relevant to their specific function(s). In particular, phenylalanine residue in position 105 (Phe105) of Alix belongs to a long loop that is unique to its Bro1 domain. Concurrently, mutation of Phe105 and surrounding residues at the tip of the loop compromise the function of Alix in HIV-1 budding without affecting its interactions with Gag or CHMP4. These studies identify a new functional determinant in the Bro1 domain of Alix.
- Laboratory of Molecular Microbiology, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: