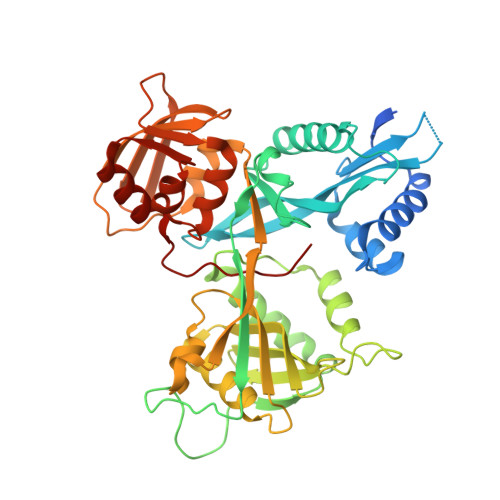
Unusual regioversatility of acetyltransferase Eis, a cause of drug resistance in XDR-TB.
Chen, W., Biswas, T., Porter, V.R., Tsodikov, O.V., Garneau-Tsodikova, S.(2011) Proc Natl Acad Sci U S A 108: 9804-9808
- PubMed: 21628583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1105379108
- Primary Citation Related Structures:
3R1K - PubMed Abstract:
The emergence of multidrug-resistant and extensively drug-resistant (XDR) tuberculosis (TB) is a serious global threat. Aminoglycoside antibiotics are used as a last resort to treat XDR-TB. Resistance to the aminoglycoside kanamycin is a hallmark of XDR-TB. Here, we reveal the function and structure of the mycobacterial protein Eis responsible for resistance to kanamycin in a significant fraction of kanamycin-resistant Mycobacterium tuberculosis clinical isolates. We demonstrate that Eis has an unprecedented ability to acetylate multiple amines of many aminoglycosides. Structural and mutagenesis studies of Eis indicate that its acetylation mechanism is enabled by a complex tripartite fold that includes two general control non-derepressible 5 (GCN5)-related N-acetyltransferase regions. An intricate negatively charged substrate-binding pocket of Eis is a potential target of new antitubercular drugs expected to overcome aminoglycoside resistance.
- Life Sciences Institute, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: