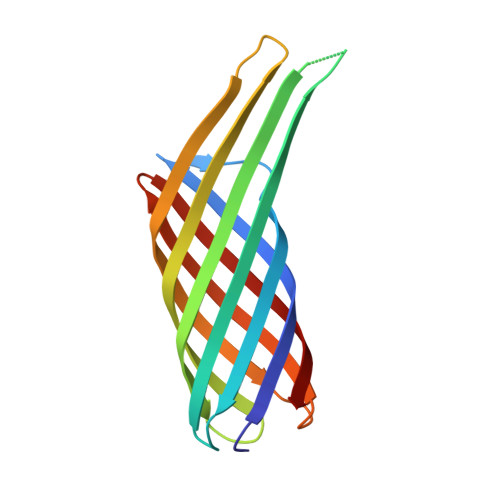Structural Insights into Ail-Mediated Adhesion in Yersinia pestis.
Yamashita, S., Lukacik, P., Barnard, T.J., Noinaj, N., Felek, S., Tsang, T.M., Krukonis, E.S., Hinnebusch, B.J., Buchanan, S.K.(2011) Structure 19: 1672-1682
- PubMed: 22078566 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.08.010
- Primary Citation Related Structures:
3QRA, 3QRC - PubMed Abstract:
Ail is an outer membrane protein from Yersinia pestis that is highly expressed in a rodent model of bubonic plague, making it a good candidate for vaccine development. Ail is important for attaching to host cells and evading host immune responses, facilitating rapid progression of a plague infection. Binding to host cells is important for injection of cytotoxic Yersinia outer proteins. To learn more about how Ail mediates adhesion, we solved two high-resolution crystal structures of Ail, with no ligand bound and in complex with a heparin analog called sucrose octasulfate. We identified multiple adhesion targets, including laminin and heparin, and showed that a 40 kDa domain of laminin called LG4-5 specifically binds to Ail. We also evaluated the contribution of laminin to delivery of Yops to HEp-2 cells. This work constitutes a structural description of how a bacterial outer membrane protein uses a multivalent approach to bind host cells.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892-8030, USA.
Organizational Affiliation: