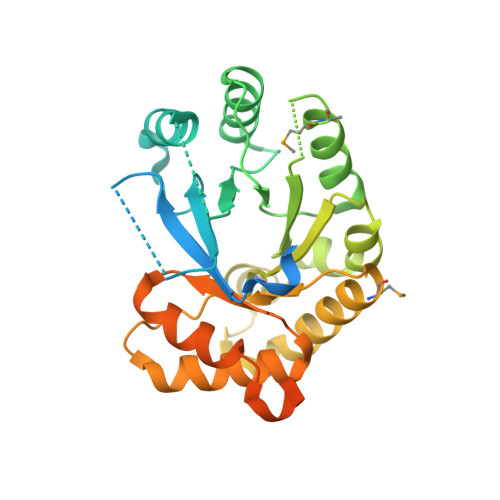
Structural insights into RipC, a putative citrate lyase beta subunit from a Yersinia pestis virulence operon
Torres, R., Chim, N., Sankaran, B., Pujol, C., Bliska, J.B., Goulding, C.W.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 2-7
- PubMed: 22232161 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111048056
- Primary Citation Related Structures:
3QLL - PubMed Abstract:
Yersinia pestis remains a threat, with outbreaks of plague occurring in rural areas and its emergence as a weapon of bioterrorism; thus, an improved understanding of its various pathogenicity pathways is warranted. The rip (required for intracellular proliferation) virulence operon is required for Y. pestis survival in interferon-γ-treated macrophages and has been implicated in lowering macrophage-produced nitric oxide levels. RipC, one of three gene products from the rip operon, is annotated as a citrate lyase β subunit. Furthermore, the Y. pestis genome lacks genes that encode citrate lyase α and γ subunits, suggesting a unique functional role of RipC in the Y. pestis rip-mediated survival pathway. Here, the 2.45 Å resolution crystal structure of RipC revealed a homotrimer in which each monomer consists of a (β/α)(8) TIM-barrel fold. Furthermore, the trimeric state was confirmed in solution by size-exclusion chromatography. Through sequence and structure comparisons with homologous proteins, it is proposed that RipC is a putative CoA- or CoA-derivative binding protein.
- Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697, USA.
Organizational Affiliation: