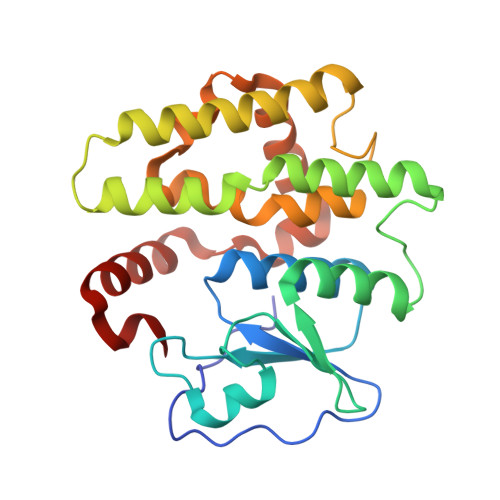
Structural insights into the dehydroascorbate reductase activity of human omega-class glutathione transferases.
Zhou, H., Brock, J., Liu, D., Board, P.G., Oakley, A.J.(2012) J Mol Biology 420: 190-203
- PubMed: 22522127 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.04.014
- Primary Citation Related Structures:
3Q18, 3Q19, 3QAG, 3VLN - PubMed Abstract:
The reduction of dehydroascorbate (DHA) to ascorbic acid (AA) is a vital cellular function. The omega-class glutathione transferases (GSTs) catalyze several reductive reactions in cellular biochemistry, including DHA reduction. In humans, two isozymes (GSTO1-1 and GSTO2-2) with significant DHA reductase (DHAR) activity are found, sharing 64% sequence identity. While the activity of GSTO2-2 is higher, it is significantly more unstable in vitro. We report the first crystal structures of human GSTO2-2, stabilized through site-directed mutagenesis and determined at 1.9 Å resolution in the presence and absence of glutathione (GSH). The structure of a human GSTO1-1 has been determined at 1.7 Å resolution in complex with the reaction product AA, which unexpectedly binds in the G-site, where the glutamyl moiety of GSH binds. The structure suggests a similar mode of ascorbate binding in GSTO2-2. This is the first time that a non-GSH-based reaction product has been observed in the G-site of any GST. AA stacks against a conserved aromatic residue, F34 (equivalent to Y34 in GSTO2-2). Mutation of Y34 to alanine in GSTO2-2 eliminates DHAR activity. From these structures and other biochemical data, we propose a mechanism of substrate binding and catalysis of DHAR activity.
- John Curtin School of Medical Research, Australian National University, Canberra, ACT 0200, Australia.
Organizational Affiliation: