Inhibitors of Ketohexokinase: Discovery of Pyrimidinopyrimidines with Specific Substitution that Complements the ATP-Binding Site.
Maryanoff, B.E., O'Neill, J.C., McComsey, D.F., Yabut, S.C., Luci, D.K., Jordan, A.D., Masucci, J.A., Jones, W.J., Abad, M.C., Gibbs, A.C., Petrounia, I.(2011) ACS Med Chem Lett 2: 538-543
- PubMed: 24900346 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml200070g
- Primary Citation Related Structures:
3Q92, 3QA2, 3QAI - PubMed Abstract:
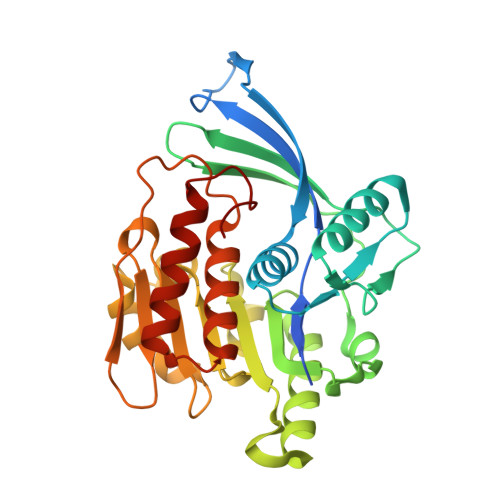
Attenuation of fructose metabolism by the inhibition of ketohexokinase (KHK; fructokinase) should reduce body weight, free fatty acids, and triglycerides, thereby offering a novel approach to treat diabetes and obesity in response to modern diets. We have identified potent, selective inhibitors of human hepatic KHK within a series of pyrimidinopyrimidines (1). For example, 8, 38, and 47 exhibited KHK IC50 values of 12, 7, and 8 nM, respectively, and also showed potent cellular KHK inhibition (IC50 < 500 nM), which relates to their intrinsic potency vs KHK and their ability to penetrate cells. X-ray cocrystal structures of KHK complexes of 3, 8, and 47 revealed the important interactions within the enzyme's adenosine 5'-triphosphate (ATP)-binding pocket.
- Johnson & Johnson Pharmaceutical Research & Development , Spring House, Pennsylvania 19477-0776, United States.
Organizational Affiliation: