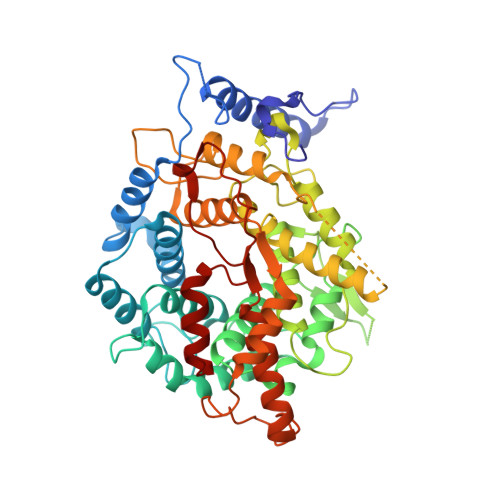
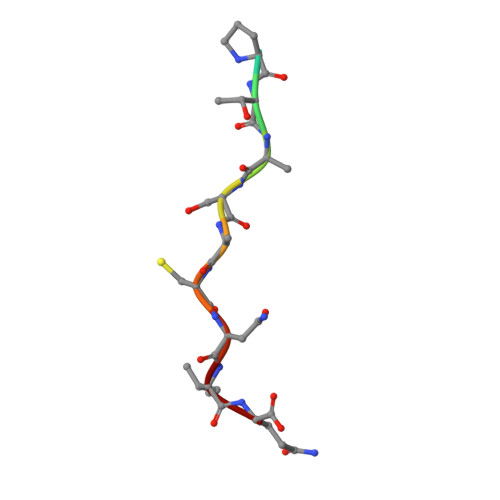
Structures of Cryptococcus neoformans Protein Farnesyltransferase Reveal Strategies for Developing Inhibitors That Target Fungal Pathogens.
Hast, M.A., Nichols, C.B., Armstrong, S.M., Kelly, S.M., Hellinga, H.W., Alspaugh, J.A., Beese, L.S.(2011) J Biological Chem 286: 35149-35162
- PubMed: 21816822 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.250506
- Primary Citation Related Structures:
3Q73, 3Q75, 3Q78, 3Q79, 3Q7A, 3Q7F, 3SFX, 3SFY - PubMed Abstract:
Cryptococcus neoformans is a fungal pathogen that causes life-threatening infections in immunocompromised individuals, including AIDS patients and transplant recipients. Few antifungals can treat C. neoformans infections, and drug resistance is increasing. Protein farnesyltransferase (FTase) catalyzes post-translational lipidation of key signal transduction proteins and is essential in C. neoformans. We present a multidisciplinary study validating C. neoformans FTase (CnFTase) as a drug target, showing that several anticancer FTase inhibitors with disparate scaffolds can inhibit C. neoformans and suggesting structure-based strategies for further optimization of these leads. Structural studies are an essential element for species-specific inhibitor development strategies by revealing similarities and differences between pathogen and host orthologs that can be exploited. We, therefore, present eight crystal structures of CnFTase that define the enzymatic reaction cycle, basis of ligand selection, and structurally divergent regions of the active site. Crystal structures of clinically important anticancer FTase inhibitors in complex with CnFTase reveal opportunities for optimization of selectivity for the fungal enzyme by modifying functional groups that interact with structurally diverse regions. A substrate-induced conformational change in CnFTase is observed as part of the reaction cycle, a feature that is mechanistically distinct from human FTase. Our combined structural and functional studies provide a framework for developing FTase inhibitors to treat invasive fungal infections.
- Department of Biochemistry, Duke University Medical Center, Durham, North Carolina 27710, USA.
Organizational Affiliation: