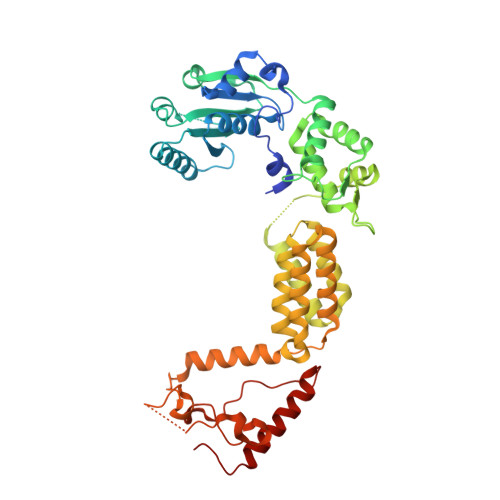
Structure and Biochemical Activities of Escherichia coli MgsA.
Page, A.N., George, N.P., Marceau, A.H., Cox, M.M., Keck, J.L.(2011) J Biological Chem 286: 12075-12085
- PubMed: 21297161 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.210187
- Primary Citation Related Structures:
3PVS - PubMed Abstract:
Bacterial "maintenance of genome stability protein A" (MgsA) and related eukaryotic enzymes play important roles in cellular responses to stalled DNA replication processes. Sequence information identifies MgsA enzymes as members of the clamp loader clade of AAA+ proteins, but structural information defining the family has been limited. Here, the x-ray crystal structure of Escherichia coli MgsA is described, revealing a homotetrameric arrangement for the protein that distinguishes it from other clamp loader clade AAA+ proteins. Each MgsA protomer is composed of three elements as follows: ATP-binding and helical lid domains (conserved among AAA+ proteins) and a tetramerization domain. Although the tetramerization domains bury the greatest amount of surface area in the MgsA oligomer, each of the domains participates in oligomerization to form a highly intertwined quaternary structure. Phosphate is bound at each AAA+ ATP-binding site, but the active sites do not appear to be in a catalytically competent conformation due to displacement of Arg finger residues. E. coli MgsA is also shown to form a complex with the single-stranded DNA-binding protein through co-purification and biochemical studies. MgsA DNA-dependent ATPase activity is inhibited by single-stranded DNA-binding protein. Together, these structural and biochemical observations provide insights into the mechanisms of MgsA family AAA+ proteins.
- Department of Biochemistry, University of Wisconsin, School of Medicine and Public Health, Madision, WI 53706, USA.
Organizational Affiliation: