Structure of the activating natural killer cell receptor NKp30 bound to its ligand B7-H6 reveals basis for tumor cell recognition in humans
Li, Y., Wang, Q., Mariuzza, R.A.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ig-like domain-containing protein DKFZp686O24166/DKFZp686I21167 | 248 | Homo sapiens | Mutation(s): 0 | 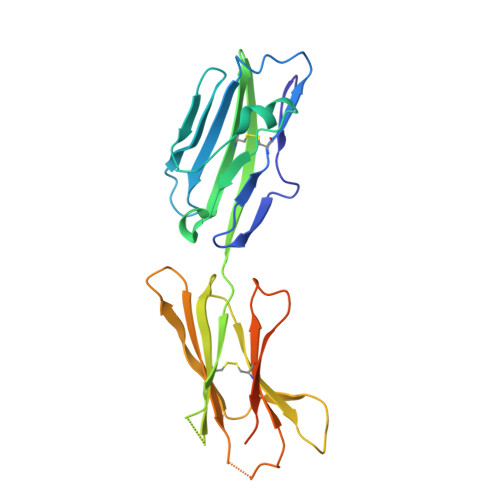 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q68D85 (Homo sapiens) Explore Q68D85 Go to UniProtKB: Q68D85 | |||||
PHAROS: Q68D85 GTEx: ENSG00000188211 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q68D85 | ||||
Glycosylation | |||||
| Glycosylation Sites: 4 | Go to GlyGen: Q68D85-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | C [auth A], D [auth A], E [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 44.3 | α = 90 |
| b = 70.6 | β = 100.4 |
| c = 53.2 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SCALEPACK | data scaling |
| PHASER | phasing |
| CNS | refinement |
| PDB_EXTRACT | data extraction |
| ADSC | data collection |
| DENZO | data reduction |