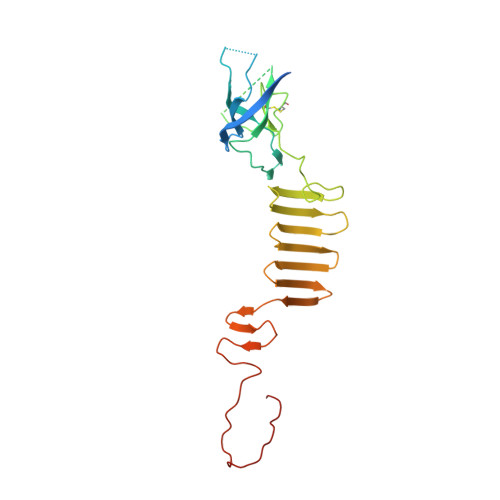
Phage pierces the host cell membrane with the iron-loaded spike.
Browning, C., Shneider, M.M., Bowman, V.D., Schwarzer, D., Leiman, P.G.(2012) Structure 20: 326-339
- PubMed: 22325780
- DOI: https://doi.org/10.1016/j.str.2011.12.009
- Primary Citation of Related Structures:
3PQH, 3PQI, 3QR7, 3QR8 - PubMed Abstract:
Bacteriophages with contractile tails and the bacterial type VI secretion system have been proposed to use a special protein to create an opening in the host cell membrane during infection. These proteins have a modular architecture but invariably contain an oligonucleotide/oligosaccharide-binding (OB-fold) domain and a long β-helical C-terminal domain, which initiates the contact with the host cell membrane. Using X-ray crystallography and electron microscopy, we report the atomic structure of the membrane-piercing proteins from bacteriophages P2 and ϕ92 and identify the residues that constitute the membrane-attacking apex. Both proteins form compact spikes with a ∼10Å diameter tip that is stabilized by a centrally positioned iron ion bound by six histidine residues. The accumulated data strongly suggest that, in the process of membrane penetration, the spikes are translocated through the lipid bilayer without undergoing major unfolding.
- Institut de Physique des Systèmes Biologiques, Laboratory of Structural Biology and Biophysics, Ecole Polytechnique Fédérale de Lausanne (EPFL), BSP-415, CH-1015, Lausanne, Switzerland.
Organizational Affiliation: