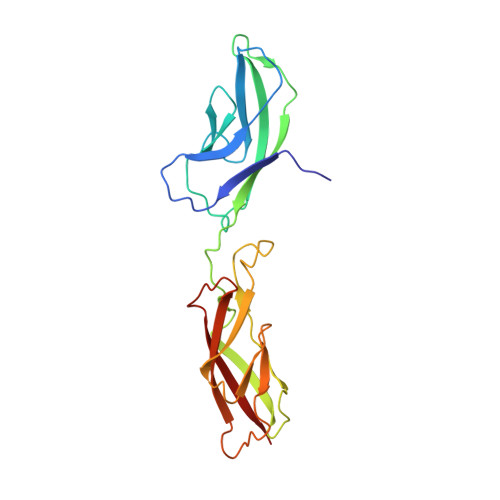
Structure and binding mechanism of vascular endothelial cadherin: a divergent classical cadherin.
Brasch, J., Harrison, O.J., Ahlsen, G., Carnally, S.M., Henderson, R.M., Honig, B., Shapiro, L.(2011) J Mol Biology 408: 57-73
- PubMed: 21269602 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.01.031
- Primary Citation Related Structures:
3PPE - PubMed Abstract:
Vascular endothelial cadherin (VE-cadherin), a divergent member of the type II classical cadherin family of cell adhesion proteins, mediates homophilic adhesion in the vascular endothelium. Previous investigations with a bacterially produced protein suggested that VE-cadherin forms cell surface trimers that bind between apposed cells to form hexamers. Here we report studies of mammalian-produced VE-cadherin ectodomains suggesting that, like other classical cadherins, VE-cadherin forms adhesive trans dimers between monomers located on opposing cell surfaces. Trimerization of the bacterially produced protein appears to be an artifact that arises from a lack of glycosylation. We also present the 2.1-Å-resolution crystal structure of the VE-cadherin EC1-2 adhesive region, which reveals homodimerization via the strand-swap mechanism common to classical cadherins. In common with type II cadherins, strand-swap binding involves two tryptophan anchor residues, but the adhesive interface resembles type I cadherins in that VE-cadherin does not form a large nonswapped hydrophobic surface. Thus, VE-cadherin is an outlier among classical cadherins, with characteristics of both type I and type II subfamilies.
- Department of Biochemistry and Molecular Biophysics, Columbia University, 701 West 168th Street, New York, NY 10032, USA.
Organizational Affiliation: