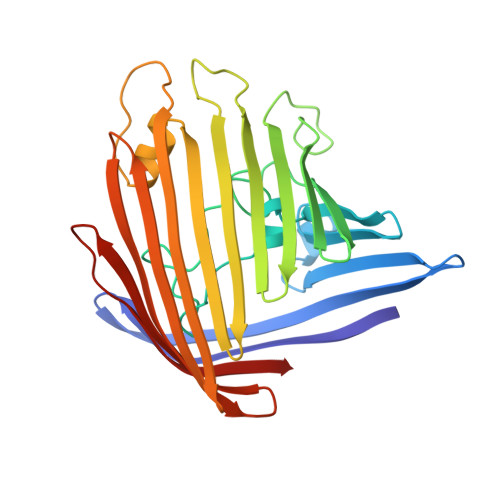
Porin conformation in the absence of calcium. Refined structure at 2.5 A resolution.
Weiss, M.S., Schulz, G.E.(1993) J Mol Biology 231: 817-824
- PubMed: 7685826 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1328
- Primary Citation Related Structures:
3POR - PubMed Abstract:
The crystal structure of porin from Rhodobacter capsulatus in the absence of divalent calcium ions has been refined to convergence at a resolution of 2.5 A using the simulated annealing refinement method. The final model consists of all 301 amino acid residues, 77 solvent molecules, one tris(hydroxymethyl)-aminomethane molecule and one unknown ligand modeled as n-octyltetraoxyethylene. A superposition with the previously described model containing three calcium ions showed structural changes at the segment 108-116 of the inner loop beta 5-beta 6, and at loops beta 8-beta 9 and beta 11-beta 12 at the extracellular side of the porin molecule. Evidence is presented that the conformational changes depend on the presence or absence of calcium ions. A possible influence on porin function is discussed.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Freiburg im Breisgau, Germany.
Organizational Affiliation: