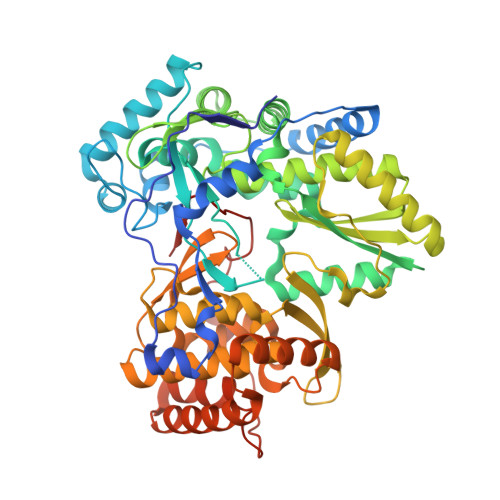
Quinolones as HCV NS5B polymerase inhibitors.
Kumar, D.V., Rai, R., Brameld, K.A., Somoza, J.R., Rajagopalan, R., Janc, J.W., Xia, Y.M., Ton, T.L., Shaghafi, M.B., Hu, H., Lehoux, I., To, N., Young, W.B., Green, M.J.(2011) Bioorg Med Chem Lett 21: 82-87
- PubMed: 21145235
- DOI: https://doi.org/10.1016/j.bmcl.2010.11.068
- Primary Citation Related Structures:
3PHE, 3UDL - PubMed Abstract:
Hepatitis C virus (HCV) infection is treated with a combination of peginterferon alfa-2a/b and ribavirin. To address the limitations of this therapy, numerous small molecule agents are in development, which act by directly affecting key steps in the viral life-cycle. Herein we describe our discovery of quinolone derivatives, novel small-molecules that inhibit NS5b polymerase, a key enzyme of the viral life-cycle. A crystal structure of a quinoline analog bound to NS5B reveals that this class of compounds binds to allosteric site-II (non-nucleoside inhibitor-site 2, NNI-2) of this protein.
- Myriad Pharmaceuticals, 305 Chipeta Way, Salt Lake City, UT 84108, USA.
Organizational Affiliation: