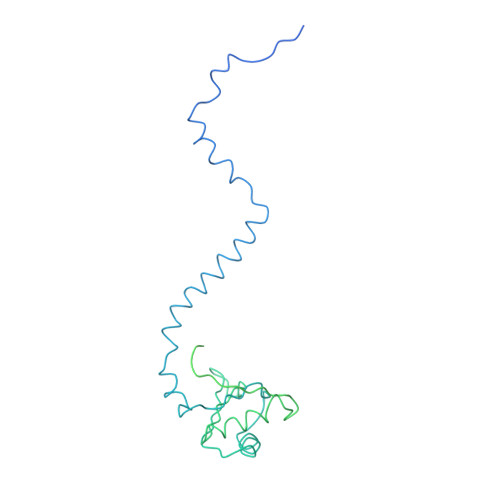
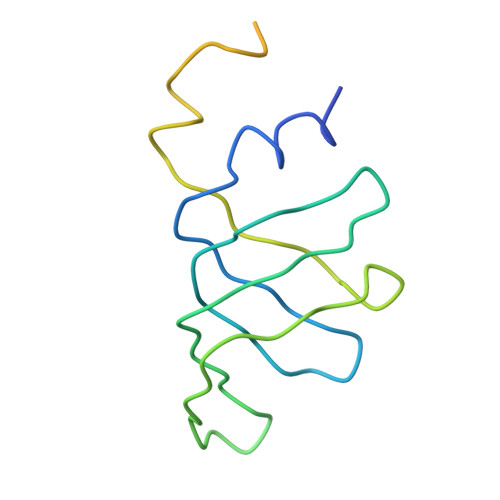
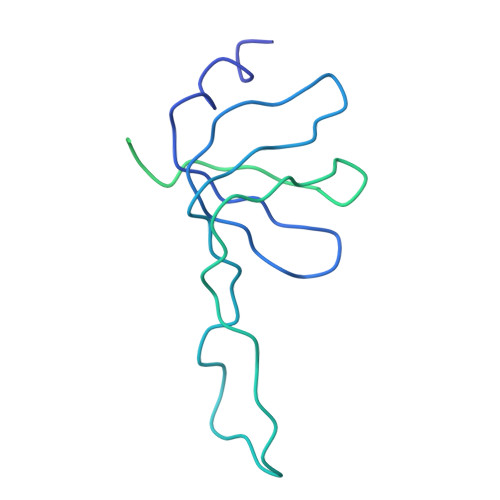
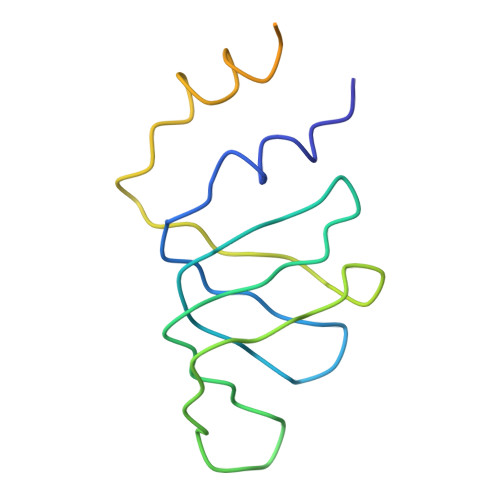
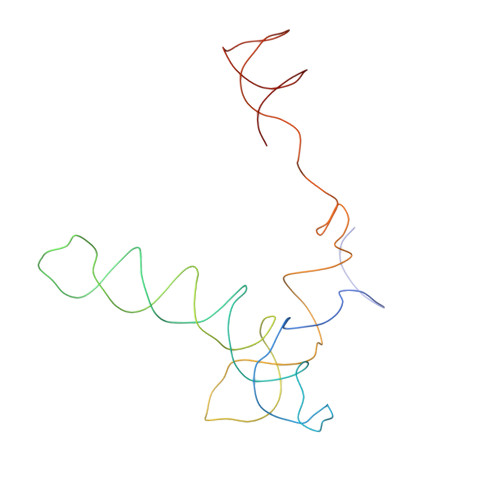
Functional organization of the Sm core in the crystal structure of human U1 snRNP.
Weber, G., Trowitzsch, S., Kastner, B., Luhrmann, R., Wahl, M.C.(2010) EMBO J 29: 4172-4184
- PubMed: 21113136 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2010.295
- Primary Citation Related Structures:
3PGW - PubMed Abstract:
U1 small nuclear ribonucleoprotein (snRNP) recognizes the 5'-splice site early during spliceosome assembly. It represents a prototype spliceosomal subunit containing a paradigmatic Sm core RNP. The crystal structure of human U1 snRNP obtained from natively purified material by in situ limited proteolysis at 4.4 Å resolution reveals how the seven Sm proteins, each recognize one nucleotide of the Sm site RNA using their Sm1 and Sm2 motifs. Proteins D1 and D2 guide the snRNA into and out of the Sm ring, and proteins F and E mediate a direct interaction between the Sm site termini. Terminal extensions of proteins D1, D2 and B/B', and extended internal loops in D2 and B/B' support a four-way RNA junction and a 3'-terminal stem-loop on opposite sides of the Sm core RNP, respectively. On a higher organizational level, the core RNP presents multiple attachment sites for the U1-specific 70K protein. The intricate, multi-layered interplay of proteins and RNA rationalizes the hierarchical assembly of U snRNPs in vitro and in vivo.
- Department of Cellular Biochemistry, Max-Planck-Institute for Biophysical Chemistry, Göttingen, Germany.
Organizational Affiliation: