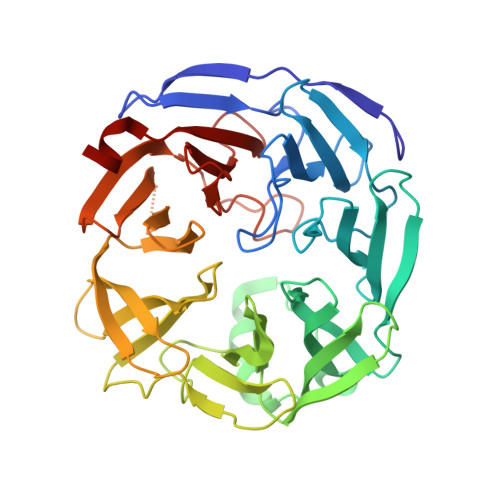
The active site of oligogalacturonate lyase provides unique insights into cytoplasmic oligogalacturonate beta-elimination.
Abbott, D.W., Gilbert, H.J., Boraston, A.B.(2010) J Biological Chem 285: 39029-39038
- PubMed: 20851883 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.153981
- Primary Citation Related Structures:
3PE7 - PubMed Abstract:
Oligogalacturonate lyases (OGLs; now also classified as pectate lyase family 22) are cytoplasmic enzymes found in pectinolytic members of Enterobacteriaceae, such as the enteropathogen Yersinia enterocolitica. OGLs utilize a β-elimination mechanism to preferentially catalyze the conversion of saturated and unsaturated digalacturonate into monogalacturonate and the 4,5-unsaturated monogalacturonate-like molecule, 5-keto-4-deoxyuronate. To provide mechanistic insights into the specificity of this enzyme activity, we have characterized the OGL from Y. enterocolitica, YeOGL, on oligogalacturonides and determined its three-dimensional x-ray structure to 1.65 Å. The model contains a Mn(2+) atom in the active site, which is coordinated by three histidines, one glutamine, and an acetate ion. The acetate mimics the binding of the uronate group of galactourono-configured substrates. These findings, in combination with enzyme kinetics and metal supplementation assays, provide a framework for modeling the active site architecture of OGL. This enzyme appears to contain a histidine for the abstraction of the α-proton in the -1 subsite, a residue that is highly conserved throughout the OGL family and represents a unique catalytic base among pectic active lyases. In addition, we present a hypothesis for an emerging relationship observed between the cellular distribution of pectate lyase folding and the distinct metal coordination chemistries of pectate lyases.
- Complex Carbohydrate Research Center, University of Georgia, Athens, Georgia 30602, USA. wabbott@ccrc.uga.edu
Organizational Affiliation: