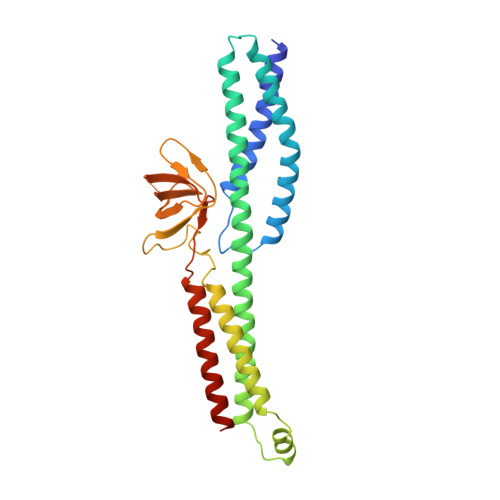
The Structure of the Plakin Domain of Plectin Reveals a Non-canonical SH3 Domain Interacting with Its Fourth Spectrin Repeat.
Ortega, E., Buey, R.M., Sonnenberg, A., de Pereda, J.M.(2011) J Biological Chem 286: 12429-12438
- PubMed: 21288893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.197467
- Primary Citation Related Structures:
3PDY, 3PE0 - PubMed Abstract:
Plectin belongs to the plakin family of cytoskeletal crosslinkers, which is part of the spectrin superfamily. Plakins contain an N-terminal conserved region, the plakin domain, which is formed by an array of spectrin repeats (SR) and a Src-homology 3 (SH3), and harbors binding sites for junctional proteins. We have combined x-ray crystallography and small angle x-ray scattering (SAXS) to elucidate the structure of the central region of the plakin domain of plectin, which corresponds to the SR3, SR4, SR5, and SH3 domains. The crystal structures of the SR3-SR4 and SR4-SR5-SH3 fragments were determined to 2.2 and 2.95 Å resolution, respectively. The SH3 of plectin presents major alterations as compared with canonical Pro-rich binding SH3 domains, suggesting that plectin does not recognize Pro-rich motifs. In addition, the SH3 binding site is partially occluded by an intramolecular contact with the SR4. Residues of this pseudo-binding site and the SR4/SH3 interface are conserved within the plakin family, suggesting that the structure of this part of the plectin molecule is similar to that of other plakins. We have created a model for the SR3-SR4-SR5-SH3 region, which agrees well with SAXS data in solution. The three SRs form a semi-flexible rod that is not altered by the presence of the SH3 domain, and it is similar to those found in spectrins. The flexibility of the plakin domain, in analogy with spectrins, might contribute to the role of plakins in maintaining the stability of tissues subject to mechanical stress.
- Instituto de Biología Molecular y Celular del Cancer, Consejo Superior de Investigaciones Científicas, University of Salamanca, Campus Unamuno, E-37007 Salamanca, Spain.
Organizational Affiliation: