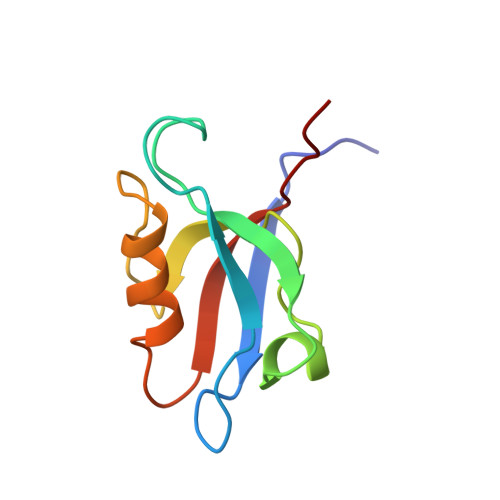
Solution structure of the PDZ2 domain from human phosphatase hPTP1E and its interactions with C-terminal peptides from the Fas receptor.
Kozlov, G., Gehring, K., Ekiel, I.(2000) Biochemistry 39: 2572-2580
- PubMed: 10704206 Search on PubMed
- DOI: https://doi.org/10.1021/bi991913c
- Primary Citation Related Structures:
3PDZ - PubMed Abstract:
The solution structure of the second PDZ domain (PDZ2) from human phosphatase hPTP1E has been determined using 2D and 3D heteronuclear NMR experiments. The binding of peptides derived from the C-terminus of the Fas receptor to PDZ2 was studied via changes in backbone peptide and protein resonances. The structure is based on a total of 1387 nonredundant experimental NMR restraints including 1261 interproton distance restraints, 45 backbone hydrogen bonds, and 81 torsion angle restraints. Analysis of 30 lowest-energy structures resulted in rmsd values of 0.41 +/- 0.09 A for backbone atoms (N, Calpha, C') and 1.08 +/- 0.10 A for all heavy atoms, excluding the disordered N- and C-termini. The hPTP1E PDZ2 structure is similar to known PDZ domain structures but contains two unique structural features. In the peptide binding domain, the first glycine of the GLGF motif is replaced by a serine. This serine appears to replace a bound water observed in PDZ crystal structures that hydrogen bonds to the bound peptide's C-terminus. The hPTP1E PDZ2 structure also contains an unusually large loop following strand beta2 and proximal to the peptide binding site. This well-ordered loop folds back against the PDZ domain and contains several residues that undergo large amide chemical shift changes upon peptide binding. Direct observation of peptide resonances demonstrates that as many as six Fas peptide residues interact with the PDZ2 domain.
- NMR Group, Pharmaceutical Biotechnology Sector and Montreal Joint Center for Structural Biology, Biotechnology Research Institute, National Research Council of Canada, Montreal, Quebec, H4P 2R2, Canada.
Organizational Affiliation: