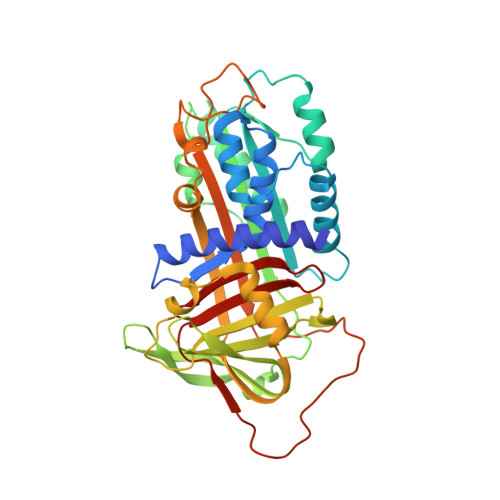
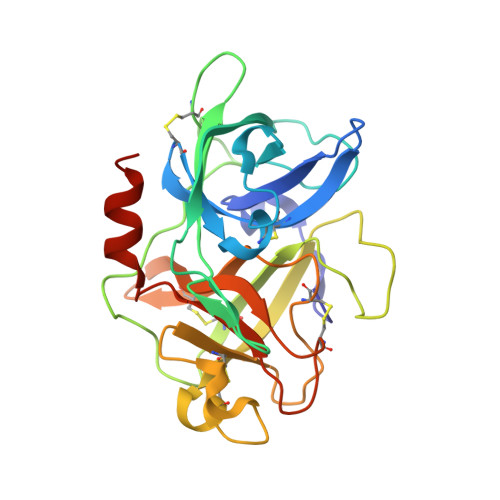
Structural basis for recognition of urokinase-type plasminogen activator by plasminogen activator inhibitor-1.
Lin, Z., Jiang, L., Yuan, C., Jensen, J.K., Zhang, X., Luo, Z., Furie, B.C., Furie, B., Andreasen, P.A., Huang, M.(2011) J Biological Chem 286: 7027-7032
- PubMed: 21199867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.204537
- Primary Citation Related Structures:
3PB1 - PubMed Abstract:
Plasminogen activator inhibitor-1 (PAI-1), together with its physiological target urokinase-type plasminogen activator (uPA), plays a pivotal role in fibrinolysis, cell migration, and tissue remodeling and is currently recognized as being among the most extensively validated biological prognostic factors in several cancer types. PAI-1 specifically and rapidly inhibits uPA and tissue-type PA (tPA). Despite extensive structural/functional studies on these two reactions, the underlying structural mechanism has remained unknown due to the technical difficulties of obtaining the relevant structures. Here, we report a strategy to generate a PAI-1·uPA(S195A) Michaelis complex and present its crystal structure at 2.3-Å resolution. In this structure, the PAI-1 reactive center loop serves as a bait to attract uPA onto the top of the PAI-1 molecule. The P4-P3' residues of the reactive center loop interact extensively with the uPA catalytic site, accounting for about two-thirds of the total contact area. Besides the active site, almost all uPA exosite loops, including the 37-, 60-, 97-, 147-, and 217-loops, are involved in the interaction with PAI-1. The uPA 37-loop makes an extensive interaction with PAI-1 β-sheet B, and the 147-loop directly contacts PAI-1 β-sheet C. Both loops are important for initial Michaelis complex formation. This study lays down a foundation for understanding the specificity of PAI-1 for uPA and tPA and provides a structural basis for further functional studies.
- State Key Laboratory of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, Chinese Academy of Sciences, Fuzhou 350002, China.
Organizational Affiliation: