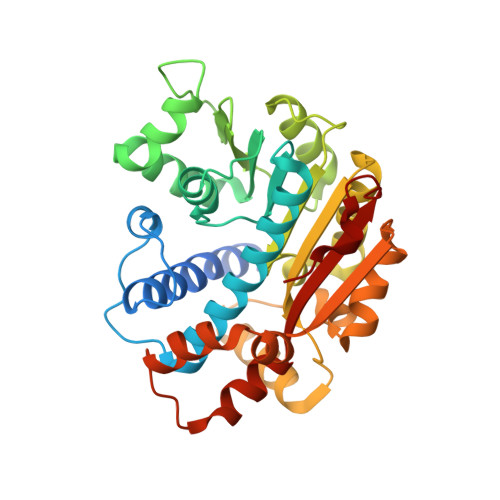
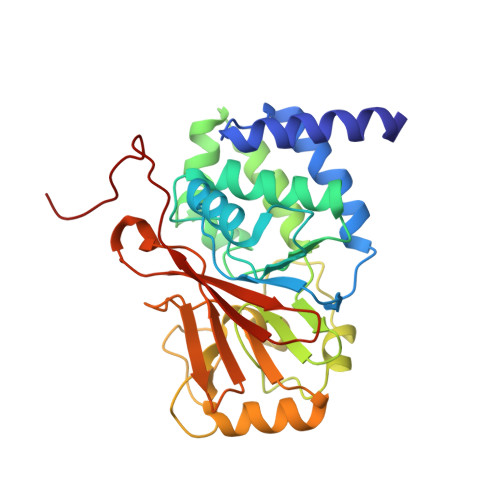
The Structural Basis for Tight Control of PP2A Methylation and Function by LCMT-1.
Stanevich, V., Jiang, L., Satyshur, K.A., Li, Y., Jeffrey, P.D., Li, Z., Menden, P., Semmelhack, M.F., Xing, Y.(2011) Mol Cell 41: 331-342
- PubMed: 21292165 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2010.12.030
- Primary Citation Related Structures:
3P71 - PubMed Abstract:
Proper formation of protein phosphatase 2A (PP2A) holoenzymes is essential for the fitness of all eukaryotic cells. Carboxyl methylation of the PP2A catalytic subunit plays a critical role in regulating holoenzyme assembly; methylation is catalyzed by PP2A-specific methyltransferase LCMT-1, an enzyme required for cell survival. We determined crystal structures of human LCMT-1 in isolation and in complex with PP2A stabilized by a cofactor mimic. The structures show that the LCMT-1 active-site pocket recognizes the carboxyl terminus of PP2A, and, interestingly, the PP2A active site makes extensive contacts to LCMT-1. We demonstrated that activation of the PP2A active site stimulates methylation, suggesting a mechanism for efficient conversion of activated PP2A into substrate-specific holoenzymes, thus minimizing unregulated phosphatase activity or formation of inactive holoenzymes. A dominant-negative LCMT-1 mutant attenuates the cell cycle without causing cell death, likely by inhibiting uncontrolled phosphatase activity. Our studies suggested mechanisms of LCMT-1 in tight control of PP2A function, important for the cell cycle and cell survival.
- McArdle Laboratory, Department of Oncology, School of Medicine and Public Health, University of Wisconsin at Madison, Madison, WI 53706, USA.
Organizational Affiliation: