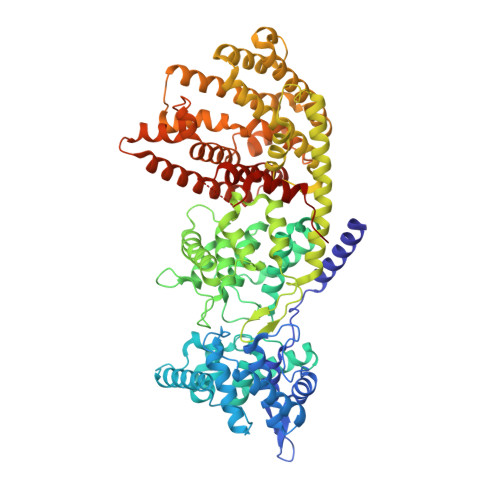
Taxadiene synthase structure and evolution of modular architecture in terpene biosynthesis.
Koksal, M., Jin, Y., Coates, R.M., Croteau, R., Christianson, D.W.(2011) Nature 469: 116-120
- PubMed: 21160477
- DOI: https://doi.org/10.1038/nature09628
- Primary Citation Related Structures:
3P5P, 3P5R - PubMed Abstract:
With more than 55,000 members identified so far in all forms of life, the family of terpene or terpenoid natural products represents the epitome of molecular biodiversity. A well-known and important member of this family is the polycyclic diterpenoid Taxol (paclitaxel), which promotes tubulin polymerization and shows remarkable efficacy in cancer chemotherapy. The first committed step of Taxol biosynthesis in the Pacific yew (Taxus brevifolia) is the cyclization of the linear isoprenoid substrate geranylgeranyl diphosphate (GGPP) to form taxa-4(5),11(12)diene, which is catalysed by taxadiene synthase. The full-length form of this diterpene cyclase contains 862 residues, but a roughly 80-residue amino-terminal transit sequence is cleaved on maturation in plastids. We now report the X-ray crystal structure of a truncation variant lacking the transit sequence and an additional 27 residues at the N terminus, hereafter designated TXS. Specifically, we have determined structures of TXS complexed with 13-aza-13,14-dihydrocopalyl diphosphate (1.82 Å resolution) and 2-fluorogeranylgeranyl diphosphate (2.25 Å resolution). The TXS structure reveals a modular assembly of three α-helical domains. The carboxy-terminal catalytic domain is a class I terpenoid cyclase, which binds and activates substrate GGPP with a three-metal ion cluster. The N-terminal domain and a third 'insertion' domain together adopt the fold of a vestigial class II terpenoid cyclase. A class II cyclase activates the isoprenoid substrate by protonation instead of ionization, and the TXS structure reveals a definitive connection between the two distinct cyclase classes in the evolution of terpenoid biosynthesis.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, 231 South 34th Street, Philadelphia, Pennsylvania 19104-6323, USA.
Organizational Affiliation: