The binding mechanism of a peptidic cyclic serine protease inhibitor
Jiang, L.G., Svane, A.S.P., Sorensen, H.P., Jensen, J.K., Hosseini, M., Chen, Z., Weydert, C., Nielsen, J.T., Christensen, A., Yuan, C., Jensen, K.J., Nielsen, N.C., Malmendal, A., Huang, M.D., Andreasen, P.A.(2011) J Mol Biology 412: 235-250
- PubMed: 21802428 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.07.028
- Primary Citation Related Structures:
3OX7, 3OY5, 3OY6 - PubMed Abstract:
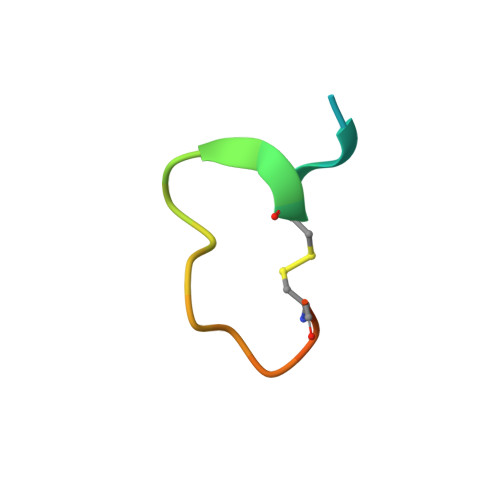
Serine proteases are classical objects for studies of catalytic and inhibitory mechanisms as well as interesting as therapeutic targets. Since small-molecule serine protease inhibitors generally suffer from specificity problems, peptidic inhibitors, isolated from phage-displayed peptide libraries, have attracted considerable attention. Here, we have investigated the mechanism of binding of peptidic inhibitors to serine protease targets. Our model is upain-1 (CSWRGLENHRMC), a disulfide-bond-constrained competitive inhibitor of human urokinase-type plasminogen activator with a noncanonical inhibitory mechanism and an unusually high specificity. Using a number of modified variants of upain-1, we characterised the upain-1-urokinase-type plasminogen activator complex using X-ray crystal structure analysis, determined a model of the peptide in solution by NMR spectroscopy, and analysed binding kinetics and thermodynamics by surface plasmon resonance and isothermal titration calorimetry. We found that upain-1 changes both main-chain conformation and side-chain orientations as it binds to the protease, in particular its Trp3 residue and the surrounding backbone. The properties of upain-1 are strongly influenced by the addition of three to four amino acids long N-terminal and C-terminal extensions to the core, disulfide-bond-constrained sequence: The C-terminal extension stabilises the solution structure compared to the core peptide alone, and the protease-bound structure of the peptide is stabilised by intrapeptide contacts between the N-terminal extension and the core peptide around Trp3. These results provide a uniquely detailed description of the binding of a peptidic protease inhibitor to its target and are of general importance in the development of peptidic inhibitors with high specificity and new inhibitory mechanisms.
- Danish-Chinese Centre for Proteases and Cancer, Aarhus University, DK-8000 Aarhus C, Denmark.
Organizational Affiliation: