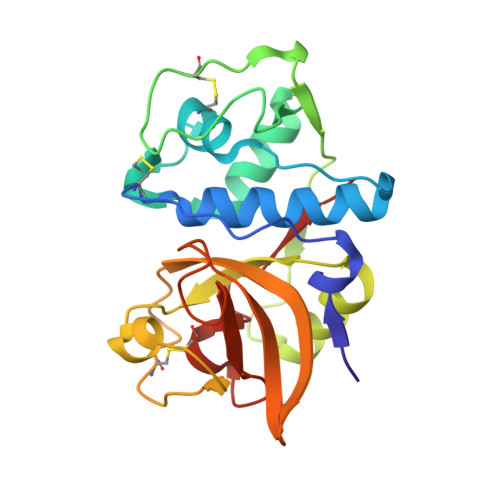
Trifluoromethylphenyl as P2 for ketoamide-based cathepsin S inhibitors.
Cai, J., Robinson, J., Belshaw, S., Everett, K., Fradera, X., van Zeeland, M., van Berkom, L., van Rijnsbergen, P., Popplestone, L., Baugh, M., Dempster, M., Bruin, J., Hamilton, W., Kinghorn, E., Westwood, P., Kerr, J., Rankovic, Z., Arbuckle, W., Bennett, D.J., Jones, P.S., Long, C., Martin, I., Uitdehaag, J.C., Meulemans, T.(2010) Bioorg Med Chem Lett 20: 6890-6894
- PubMed: 21030256 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2010.10.012
- Primary Citation Related Structures:
3OVX, 3OVZ - PubMed Abstract:
The trifluoromethylphenyl P2 motif from previously reported heteroarylnitrile series has been successfully applied for the design and synthesis of highly potent novel ketoamide-based cathepsin S inhibitors. The key in this process is the change of the torsion angle between the P2 phenyl ring and the attached secondary amide by adding a small Cl, F, or Me group at the 2-position.
- Merck Research Laboratories, MSD, Newhouse, Lanarkshire, United Kingdom. jiaqiang.cai@merck.com
Organizational Affiliation: