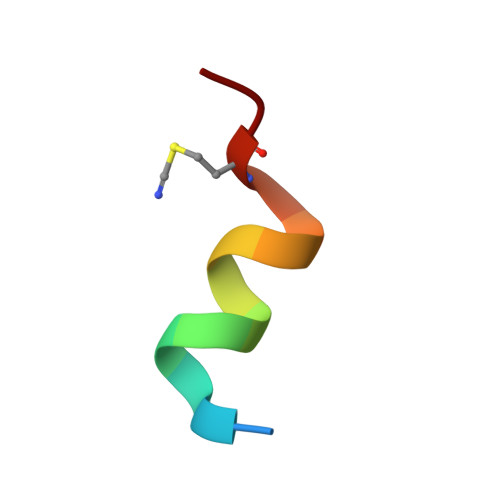
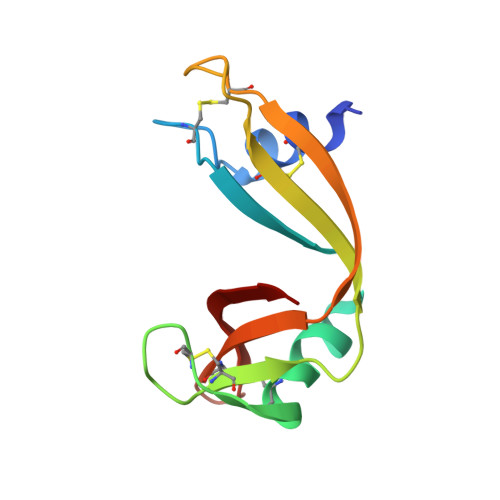
Nitrile bonds as infrared probes of electrostatics in ribonuclease S.
Fafarman, A.T., Boxer, S.G.(2010) J Phys Chem B 114: 13536-13544
- PubMed: 20883003 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jp106406p
- Primary Citation Related Structures:
3OQY, 3OQZ, 3OR0 - PubMed Abstract:
Three different nitrile-containing amino acids, p-cyanophenylalanine, m-cyanophenylalanine, and S-cyanohomocysteine, have been introduced near the active site of the semisynthetic enzyme ribonuclease S (RNase S) to serve as probes of electrostatic fields. Vibrational Stark spectra, measured directly on the probe-modified proteins, confirm the predominance of the linear Stark tuning rate in describing the sensitivity of the nitrile stretch to external electric fields, a necessary property for interpreting observed frequency shifts as a quantitative measure of local electric fields that can be compared with simulations. The X-ray structures of these nitrile-modified RNase variants and enzymatic assays demonstrate minimal perturbation to the structure and function, respectively, by the probes and provide a context for understanding the influence of the environment on the nitrile stretching frequency. We examine the ability of simulation techniques to recapitulate the spectroscopic properties of these nitriles as a means to directly test a computational electrostatic model for proteins, specifically that in the ubiquitous Amber-99 force field. Although qualitative agreement between theory and experiment is observed for the largest shifts, substantial discrepancies are observed in some cases, highlighting the ongoing need for experimental metrics to inform the development of theoretical models of electrostatic fields in proteins.
- Department of Chemistry, Stanford University, Stanford, California 94305-5080, USA.
Organizational Affiliation: