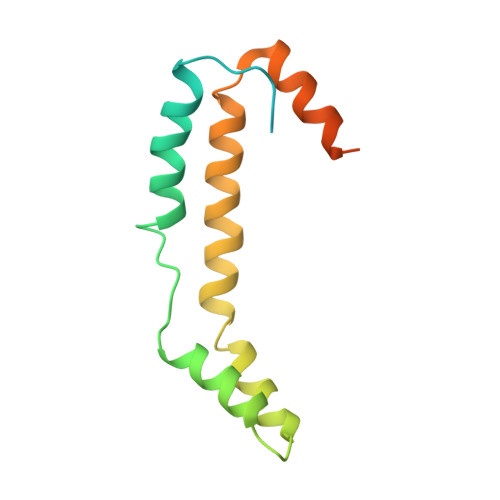
The crystal structure Escherichia coli Spy.
Kwon, E., Kim, D.Y., Gross, C.A., Gross, J.D., Kim, K.K.(2010) Protein Sci 19: 2252-2259
- PubMed: 20799348
- DOI: https://doi.org/10.1002/pro.489
- Primary Citation Related Structures:
3OEO - PubMed Abstract:
Escherichia coli spheroplast protein y (EcSpy) is a small periplasmic protein that is homologous with CpxP, an inhibitor of the extracytoplasmic stress response. Stress conditions such as spheroplast formation induce the expression of Spy via the Cpx or the Bae two-component systems in E. coli, though the function of Spy is unknown. Here, we report the crystal structure of EcSpy, which reveals a long kinked hairpin-like structure of four α-helices that form an antiparallel dimer. The dimer contains a curved oval shape with a highly positively charged concave surface that may function as a ligand binding site. Sequence analysis reveals that Spy is highly conserved over the Enterobacteriaceae family. Notably, three conserved regions that contain identical residues and two LTxxQ motifs are placed at the horizontal end of the dimer structure, stabilizing the overall fold. CpxP also contains the conserved sequence motifs and has a predicted secondary structure similar to Spy, suggesting that Spy and CpxP likely share the same fold.
- Department of Molecular Cell Biology, Samsung Biomedical Research Institute, Sungkyunkwan University School of Medicine, Suwon, Korea.
Organizational Affiliation: