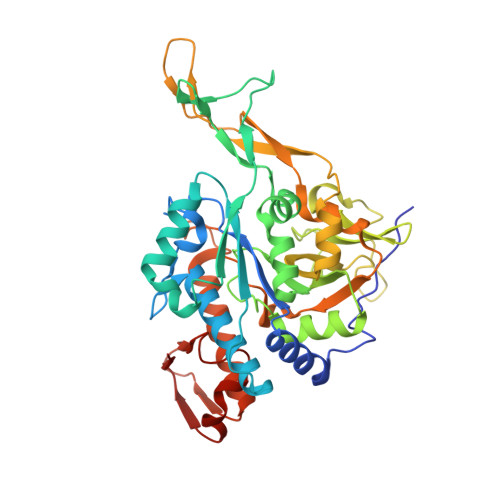
Structure of uridine diphosphate N-acetylglucosamine pyrophosphorylase from Entamoeba histolytica.
Edwards, T.E., Gardberg, A.S., Phan, I.Q., Zhang, Y., Staker, B.L., Myler, P.J., Lorimer, D.D.(2015) Acta Crystallogr Sect F Struct Biol Cryst Commun 71: 560-565
- PubMed: 25945709 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X1500179X
- Primary Citation Related Structures:
3OC9 - PubMed Abstract:
Uridine diphosphate N-acetylglucosamine pyrophosphorylase (UAP) catalyzes the final step in the synthesis of UDP-GlcNAc, which is involved in cell-wall biogenesis in plants and fungi and in protein glycosylation. Small-molecule inhibitors have been developed against UAP from Trypanosoma brucei that target an allosteric pocket to provide selectivity over the human enzyme. A 1.8 Å resolution crystal structure was determined of UAP from Entamoeba histolytica, an anaerobic parasitic protozoan that causes amoebic dysentery. Although E. histolytica UAP exhibits the same three-domain global architecture as other UAPs, it appears to lack three α-helices at the N-terminus and contains two amino acids in the allosteric pocket that make it appear more like the enzyme from the human host than that from the other parasite T. brucei. Thus, allosteric inhibitors of T. brucei UAP are unlikely to target Entamoeba UAPs.
- Seattle Structural Genomics Center for Infectious Disease, USA.
Organizational Affiliation: