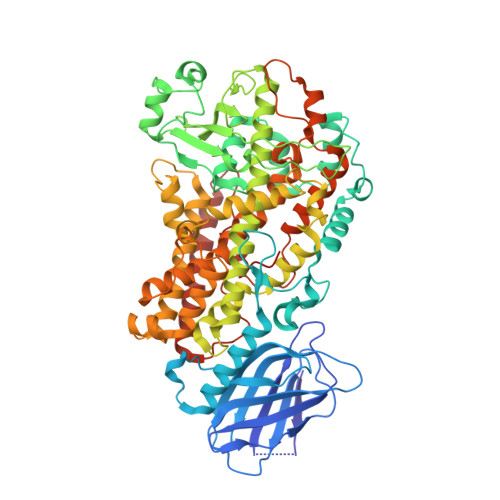
The structure of human 5-lipoxygenase.
Gilbert, N.C., Bartlett, S.G., Waight, M.T., Neau, D.B., Boeglin, W.E., Brash, A.R., Newcomer, M.E.(2011) Science 331: 217-219
- PubMed: 21233389
- DOI: https://doi.org/10.1126/science.1197203
- Primary Citation Related Structures:
3O8Y - PubMed Abstract:
The synthesis of both proinflammatory leukotrienes and anti-inflammatory lipoxins requires the enzyme 5-lipoxygenase (5-LOX). 5-LOX activity is short-lived, apparently in part because of an intrinsic instability of the enzyme. We identified a 5-LOX-specific destabilizing sequence that is involved in orienting the carboxyl terminus, which binds the catalytic iron. Here, we report the crystal structure at 2.4 angstrom resolution of human 5-LOX stabilized by replacement of this sequence.
- Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA.
Organizational Affiliation: