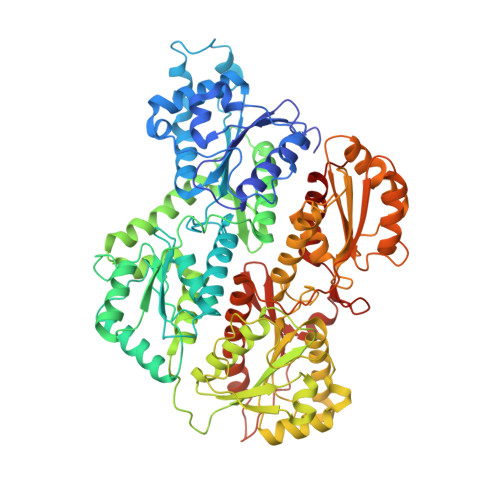
The Crystal Structures of Eukaryotic Phosphofructokinases from Baker's Yeast and Rabbit Skeletal Muscle.
Banaszak, K., Mechin, I., Obmolova, G., Oldham, M., Chang, S.H., Ruiz, T., Radermacher, M., Kopperschlager, G., Rypniewski, W.(2011) J Mol Biology 407: 284-297
- PubMed: 21241708 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.01.019
- Primary Citation Related Structures:
3O8L, 3O8N, 3O8O - PubMed Abstract:
Phosphofructokinase 1 (PFK) is a multisubunit allosteric enzyme that catalyzes the principal regulatory step in glycolysis-the phosphorylation of fructose 6-phosphate to fructose 1,6-bisphosphate by ATP. The activity of eukaryotic PFK is modulated by a number of effectors in response to the cell's needs for energy and building blocks for biosynthesis. The crystal structures of eukaryotic PFKs-from Saccharomyces cerevisiae and rabbit skeletal muscle-demonstrate how successive gene duplications and fusion are reflected in the protein structure and how they allowed the evolution of new functionalities. The basic framework inherited from prokaryotes is conserved, and additional levels of structural and functional complexity have evolved around it. Analysis of protein-ligand complexes has shown how PFK is activated by fructose 2,6-bisphosphate (a powerful PFK effector found only in eukaryotes) and reveals a novel nucleotide binding site. Crystallographic results have been used as the basis for structure-based effector design.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan, Poland.
Organizational Affiliation: