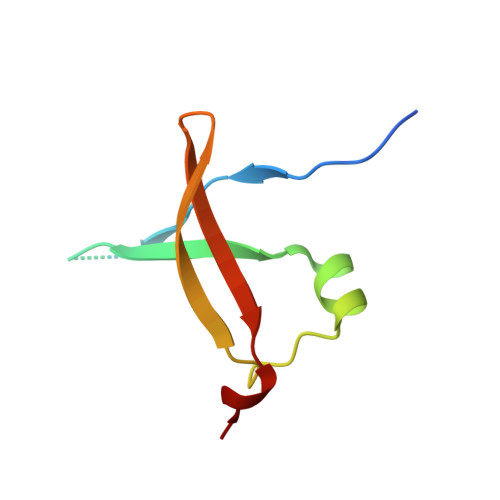
C68 from the Sulfolobus islandicus plasmid-virus pSSVx is a novel member of the AbrB-like transcription factor family.
Contursi, P., D'Ambrosio, K., Pirone, L., Pedone, E., Aucelli, T., She, Q., De Simone, G., Bartolucci, S.(2011) Biochem J 435: 157-166
- PubMed: 21208189 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20101334
- Primary Citation Related Structures:
3O27 - PubMed Abstract:
The genetic element pSSVx from Sulfolobus islandicus, strain REY15/4, is a hybrid between a plasmid and a fusellovirus. This plasmid-virus hybrid infects several species of the hyperthermophilic acidophilic crenarchaeon Sulfolobus. The open reading frame orfc68 of pSSVx encodes a 7.7 kDa protein that does not show significant sequence homology with any protein with known three-dimensional structure. EMSA (electrophoretic mobility-shift assay) experiments, DNA footprinting and CD analyses indicate that recombinant C68, purified from Escherichia coli, binds to two different operator sites that are located upstream of its own promoter. The three-dimensional structure, solved by a single-wavelength anomalous diffraction experiment on a selenomethionine derivative, shows that the protein assumes a swapped-hairpin fold, which is a distinctive fold associated with a family of prokaryotic transcription factors, such as AbrB from Bacillus subtilis. Nevertheless, C68 constitutes a novel representative of this family because it shows several peculiar structural and functional features.
- Dipartimento di Biologia Strutturale e Funzionale, Università degli Studi di Napoli Federico II, Complesso Universitario Monte S. Angelo, Via Cinthia, Naples, Italy. contursi@unina.it
Organizational Affiliation: