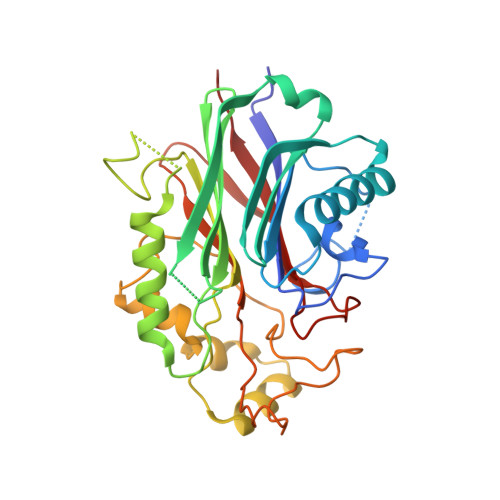
Structural basis for phosphoinositide substrate recognition, catalysis, and membrane interactions in human inositol polyphosphate 5-phosphatases
Tresaugues, L., Silvander, C., Flodin, S., Welin, M., Nyman, T., Graslund, S., Hammarstrom, M., Berglund, H., Nordlund, P.(2014) Structure 22: 744-755
- PubMed: 24704254 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2014.01.013
- Primary Citation Related Structures:
3MTC, 3N9V, 3NR8, 4CML, 4CMN - PubMed Abstract:
SHIP2, OCRL, and INPP5B belong to inositol polyphosphate 5-phophatase subfamilies involved in insulin regulation and Lowes syndrome. The structural basis for membrane recognition, substrate specificity, and regulation of inositol polyphosphate 5-phophatases is still poorly understood. We determined the crystal structures of human SHIP2, OCRL, and INPP5B, the latter in complex with phosphoinositide substrate analogs, which revealed a membrane interaction patch likely to assist in sequestering substrates from the lipid bilayer. Residues recognizing the 1-phosphate of the substrates are highly conserved among human family members, suggesting similar substrate binding modes. However, 3- and 4-phosphate recognition varies and determines individual substrate specificity profiles. The high conservation of the environment of the scissile 5-phosphate suggests a common reaction geometry for all members of the human 5-phosphatase family.
- Structural Genomics Consortium, Karolinska Institutet, 17177 Stockholm, Sweden.
Organizational Affiliation: