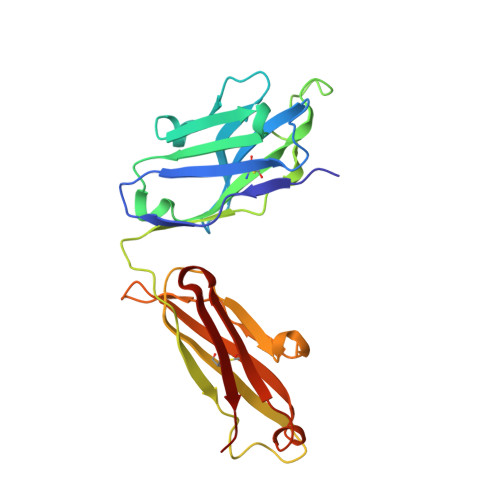
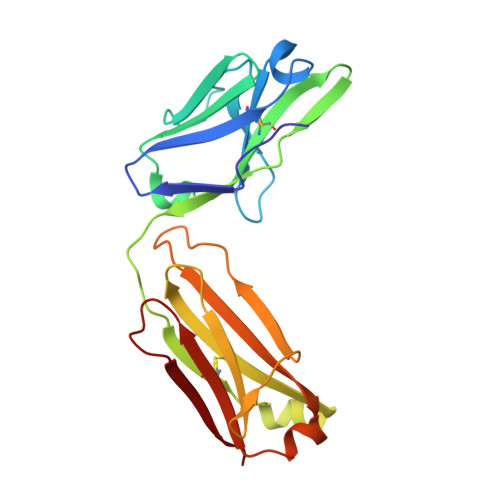
A reverse binding motif that contributes to specific protease inhibition by antibodies.
Schneider, E.L., Lee, M.S., Baharuddin, A., Goetz, D.H., Farady, C.J., Ward, M., Wang, C.I., Craik, C.S.(2012) J Mol Biology 415: 699-715
- PubMed: 22154938 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.11.036
- Primary Citation Related Structures:
3NPS, 3SO3 - PubMed Abstract:
The type II transmembrane serine protease family consists of 18 closely related serine proteases that are implicated in multiple functions. To identify selective, inhibitory antibodies against one particular type II transmembrane serine protease, matriptase [MT-SP1 (membrane-type serine protease 1)], a phage display library was created with a natural repertoire of Fabs [fragment antigen binding (Fab)] from human naïve B cells. Fab A11 was identified with a 720 pM inhibition constant and high specificity for matriptase over other trypsin-fold serine proteases. A Trichoderma reesei system expressed A11 with a yield of ∼200 mg/L. The crystal structure of A11 in complex with matriptase has been determined and compared to the crystal structure of another antibody inhibitor (S4) in complex with matriptase. Previously discovered from a synthetic single-chain variable fragment library, S4 is also a highly selective and potent matriptase inhibitor. The crystal structures of the A11/matriptase and S4/matriptase complexes were solved to 2.1 Å and 1.5 Å, respectively. Although these antibodies, discovered from separate libraries, interact differently with the protease surface loops for their specificity, the structures reveal a similar novel mechanism of protease inhibition. Through the insertion of the H3 variable loop in a reverse orientation at the substrate-binding pocket, these antibodies bury a large surface area for potent inhibition and avoid proteolytic inactivation. This discovery highlights the critical role that the antibody scaffold plays in positioning loops to bind and inhibit protease function in a highly selective manner. Additionally, Fab A11 is a fully human antibody that specifically inhibits matriptase over other closely related proteases, suggesting that this approach could be useful for clinical applications.
- Department of Pharmaceutical Chemistry, University of California, San Francisco, Genentech Hall, San Francisco, CA 94143-2280, USA.
Organizational Affiliation: